Etv transcription factors functionally diverge from their upstream FGF signaling in lens development
- PMID: 32043969
- PMCID: PMC7069720
- DOI: 10.7554/eLife.51915
Etv transcription factors functionally diverge from their upstream FGF signaling in lens development
Abstract
The signal regulated transcription factors (SRTFs) control the ultimate transcriptional output of signaling pathways. Here, we examined a family of FGF-induced SRTFs - Etv1, Etv 4, and Etv 5 - in murine lens development. Contrary to FGF receptor mutants that displayed loss of ERK signaling and defective cell differentiation, Etv deficiency augmented ERK phosphorylation without disrupting the normal lens fiber gene expression. Instead, the transitional zone for lens differentiation was shifted anteriorly as a result of reduced Jag1-Notch signaling. We also showed that Etv proteins suppresses mTOR activity by promoting Tsc2 expression, which is necessary for the nuclei clearance in mature lens. These results revealed the functional divergence between Etv and FGF in lens development, demonstrating that these SRTFs can operate outside the confine of their upstream signaling.
Keywords: ETS; FGF; Notch; PEA3; developmental biology; lens development; mTOR; mouse.
Plain language summary
Many cells contain proteins known as signal-induced transcription factors, which are poised to receive messages from the environment and then react by activating genes required for the cell to respond appropriately. It is commonly thought that these transcription factors faithfully follow the instructions they receive from the external signal: for instance, if the message was to encourage the cell to grow, the transcription factors would switch on growth-related genes. As the eyes of mice and other mammals develop, a signal known as FGF is required for certain cells to specialize into lens fiber cells: these long, thin, transparent cells form the bulk of the lens, the structure that allows focused vision. Previous studies suggest that FGF activates three transcription factors known as Etv1, Etv4 and Etv5, but their precise roles in the development of the lens has remained unclear. Here, Garg, Hannan, Wang et al. confirm that FGF signaling does indeed activate all three proteins. However, mutant mice that lacked Etv1, Etv4 and Etv5 still created lens fiber cells, suggesting that the transcription factors are largely unnecessary for lens fiber cells formation. Instead, the Etv proteins participated in a cascade of molecular events involving a protein called Notch; as a result, if the transcription factors were absent, the lens fiber cells formed prematurely. In addition, deactivating Etv1, Etv4 and Etv5 also promoted the activity of a protein which interfered with the removal of internal cell compartments, a process required for lens fiber cells to mature properly. These findings reveal that the roles of Etv1, Etv4 and Etv5 deviate from and even oppose FGF signaling in the lenses of mice. Transcription factors control the ultimate fate of a cell, and there is therefore increased interest in targeting them for therapy. The work by Garg, Hannan, Wang et al. reveals an unexpected complexity in how these proteins respond to upstream signals, highlighting the importance of further dissecting these relationships.
© 2020, Garg et al.
Conflict of interest statement
AG, AH, QW, NM, JZ, HL, SY, YM, XZ No competing interests declared
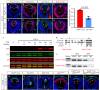
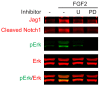
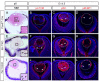
Figures

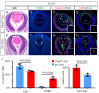

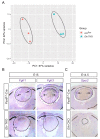

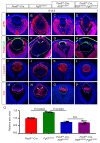
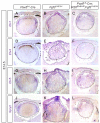

References
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

