Identification of Potentially Therapeutic Target Genes of Hepatocellular Carcinoma
- PMID: 32046048
- PMCID: PMC7037431
- DOI: 10.3390/ijerph17031053
Identification of Potentially Therapeutic Target Genes of Hepatocellular Carcinoma
Abstract
Background: Hepatocellular carcinoma (HCC) is a major threat to public health. However, few effective therapeutic strategies exist. We aimed to identify potentially therapeutic target genes of HCC by analyzing three gene expression profiles.
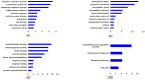
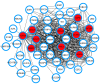
Methods: The gene expression profiles were analyzed with GEO2R, an interactive web tool for gene differential expression analysis, to identify common differentially expressed genes (DEGs). Functional enrichment analyses were then conducted followed by a protein-protein interaction (PPI) network construction with the common DEGs. The PPI network was employed to identify hub genes, and the expression level of the hub genes was validated via data mining the Oncomine database. Survival analysis was carried out to assess the prognosis of hub genes in HCC patients.
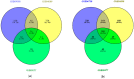
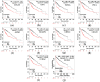
Results: A total of 51 common up-regulated DEGs and 201 down-regulated DEGs were obtained after gene differential expression analysis of the profiles. Functional enrichment analyses indicated that these common DEGs are linked to a series of cancer events. We finally identified 10 hub genes, six of which (OIP5, ASPM, NUSAP1, UBE2C, CCNA2, and KIF20A) are reported as novel HCC hub genes. Data mining the Oncomine database validated that the hub genes have a significant high level of expression in HCC samples compared normal samples (t-test, p < 0.05). Survival analysis indicated that overexpression of the hub genes is associated with a significant reduction (p < 0.05) in survival time in HCC patients.
Conclusions: We identified six novel HCC hub genes that might be therapeutic targets for the development of drugs for some HCC patients.
Keywords: MCC algorithm; PPI network; gene expression profile; hepatocellular carcinoma; hub gene.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript; or in the decision to publish the results.
Figures

References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

