The Medicinal Chemistry in the Era of Machines and Automation: Recent Advances in Continuous Flow Technology
- PMID: 32049517
- PMCID: PMC7997576
- DOI: 10.1021/acs.jmedchem.9b01956
The Medicinal Chemistry in the Era of Machines and Automation: Recent Advances in Continuous Flow Technology
Abstract
Medicinal chemistry plays a fundamental and underlying role in chemical biology, pharmacology, and medicine to discover safe and efficacious drugs. Small molecule medicinal chemistry relies on iterative learning cycles composed of compound design, synthesis, testing, and data analysis to provide new chemical probes and lead compounds for novel and druggable targets. Using traditional approaches, the time from hypothesis to obtaining the results can be protracted, thus limiting the number of compounds that can be advanced into clinical studies. This challenge can be tackled with the recourse of enabling technologies that are showing great potential in improving the drug discovery process. In this Perspective, we highlight recent developments toward innovative medicinal chemistry strategies based on continuous flow systems coupled with automation and bioassays. After a discussion of the aims and concepts, we describe equipment and representative examples of automated flow systems and end-to-end prototypes realized to expedite medicinal chemistry discovery cycles.
Conflict of interest statement
The authors declare no competing financial interest.
Figures
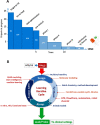
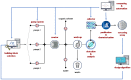





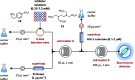

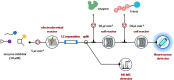



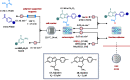
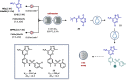
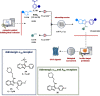

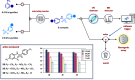



References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Chemical Information

