CRISPR-Mediated Protein Tagging with Nanoluciferase to Investigate Native Chemokine Receptor Function and Conformational Changes
- PMID: 32053779
- PMCID: PMC7242902
- DOI: 10.1016/j.chembiol.2020.01.010
CRISPR-Mediated Protein Tagging with Nanoluciferase to Investigate Native Chemokine Receptor Function and Conformational Changes
Abstract
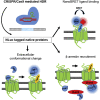
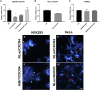
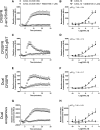
G protein-coupled receptors are a major class of membrane receptors that mediate physiological and pathophysiological cellular signaling. Many aspects of receptor activation and signaling can be investigated using genetically encoded luminescent fusion proteins. However, the use of these biosensors in live cell systems requires the exogenous expression of the tagged protein of interest. To maintain the normal cellular context here we use CRISPR/Cas9-mediated homology-directed repair to insert luminescent tags into the endogenous genome. Using NanoLuc and bioluminescence resonance energy transfer we demonstrate fluorescent ligand binding at genome-edited chemokine receptors. We also demonstrate that split-NanoLuc complementation can be used to investigate conformational changes and internalization of CXCR4 and that recruitment of β-arrestin2 to CXCR4 can be monitored when both proteins are natively expressed. These results show that genetically encoded luminescent biosensors can be used to investigate numerous aspects of receptor function at native expression levels.
Keywords: ACKR3; CXCR4; G protein-coupled receptor; NanoBRET; NanoBiT; NanoLuc; endogenous.; fluorescent ligands; ligand binding; β-arrestin2.
Copyright © 2020 The Authors. Published by Elsevier Ltd.. All rights reserved.
Conflict of interest statement
Declaration of Interests K.D.G.P. receives funding from Promega, BMG Labtech, and Dimerix as Australian Research Council Linkage Grant participating organizations. These participating organizations played no role in any aspect of the conception or design of the research, collection, analysis, and interpretation of results, or writing and editing of the manuscript. K.D.G.P. is Chief Scientific Advisor of Dimerix, of which he maintains a shareholding. The authors declare no other competing interests.
Figures




Comment in
-
Playing Tag with Your Favorite GPCR Using CRISPR.Cell Chem Biol. 2020 Jun 18;27(6):642-644. doi: 10.1016/j.chembiol.2020.06.001. Cell Chem Biol. 2020. PMID: 32559501
References
-
- Bockaert J., Fagni L., Dumuis A., Marin P. GPCR interacting proteins (GIP) Pharmacol. Ther. 2004;103:203–221. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

