CpG-ODN-mediated TLR9 innate immune signalling and calcium dyshomeostasis converge on the NFκB inhibitory protein IκBβ to drive IL1α and IL1β expression
- PMID: 32064589
- PMCID: PMC7160659
- DOI: 10.1111/imm.13182
CpG-ODN-mediated TLR9 innate immune signalling and calcium dyshomeostasis converge on the NFκB inhibitory protein IκBβ to drive IL1α and IL1β expression
Abstract
Sterile inflammation contributes to many pathological states associated with mitochondrial injury. Mitochondrial injury disrupts calcium homeostasis and results in the release of CpG-rich mitochondrial DNA. The role of CpG-stimulated TLR9 innate immune signalling and sterile inflammation is well studied; however, how calcium dyshomeostasis affects this signalling is unknown. Therefore, we interrogated the relationship beτween intracellular calcium and CpG-induced TLR9 signalling in murine macrophages. We found that CpG-ODN-induced NFκB-dependent IL1α and IL1β expression was significantly attenuated by both calcium chelation and calcineurin inhibition, a finding mediated by inhibition of degradation of the NFκB inhibitory protein IκBβ. In contrast, calcium ionophore exposure increased CpG-induced IκBβ degradation and IL1α and IL1β expression. These results demonstrate that through its effect on IκBβ degradation, increased intracellular Ca2+ drives a pro-inflammatory TLR9-mediated innate immune response. These results have implications for the study of innate immune signalling downstream of mitochondrial stress and injury.
Keywords: cytokines; inflammation; monocytes/macrophages; rodent; transcription factors; transgenic/knockout mice.
© 2020 John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare no potential financial or ethical conflicts of interest.
Figures
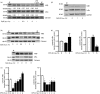
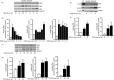



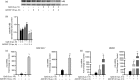
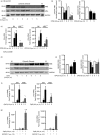
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

