The kinase polypharmacology landscape of clinical PARP inhibitors
- PMID: 32066817
- PMCID: PMC7026418
- DOI: 10.1038/s41598-020-59074-4
The kinase polypharmacology landscape of clinical PARP inhibitors
Abstract
Polypharmacology plays an important role in defining response and adverse effects of drugs. For some mechanisms, experimentally mapping polypharmacology is commonplace, although this is typically done within the same protein class. Four PARP inhibitors have been approved by the FDA as cancer therapeutics, yet a precise mechanistic rationale to guide clinicians on which to choose for a particular patient is lacking. The four drugs have largely similar PARP family inhibition profiles, but several differences at the molecular and clinical level have been reported that remain poorly understood. Here, we report the first comprehensive characterization of the off-target kinase landscape of four FDA-approved PARP drugs. We demonstrate that all four PARP inhibitors have a unique polypharmacological profile across the kinome. Niraparib and rucaparib inhibit DYRK1s, CDK16 and PIM3 at clinically achievable, submicromolar concentrations. These kinases represent the most potently inhibited off-targets of PARP inhibitors identified to date and should be investigated further to clarify their potential implications for efficacy and safety in the clinic. Moreover, broad kinome profiling is recommended for the development of PARP inhibitors as PARP-kinase polypharmacology could potentially be exploited to modulate efficacy and side-effect profiles.
Conflict of interest statement
A.A.A., M.A., U. B., P.A.C., P.W. and B.A.-L. are/were employees of The Institute of Cancer Research (ICR), which has a commercial interest in a range of drug targets, including protein kinases and are involved in a patent on the use of PARP inhibitors in BRCA-mutated cancers which results in commercial income. The ICR operates a Rewards to Inventors scheme whereby employees of the ICR may receive financial benefit following commercial licensing of a project. P.W. is a consultant/scientific advisory board member for Nextech Invest Ltd, Storm Therapeutics, Astex Pharmaceuticals, Nuevolution and CV6 and holds stock in Chroma Therapeutics, NextInvest and Storm Therapeutics; he is also a Non-Executive Director of Storm Therapeutics and the Royal Marsden NHS Trust and a Director of the non-profit Chemical Probes Portal. B.A.-L. is/was a consultant/scientific advisory board member for GSK, Open Targets, Astex Pharmaceuticals, Astellas Pharma and is an ex-employee of Inpharmatica Ltd. A.A.A., B.A.-L. and P.W. have been instrumental in the creation/development of canSAR and Probe Miner. B.A.-L was instrumental in the creation of ChEMBL.
Figures
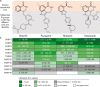



References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

