DNA nanotechnology assisted nanopore-based analysis
- PMID: 32083656
- PMCID: PMC7102975
- DOI: 10.1093/nar/gkaa095
DNA nanotechnology assisted nanopore-based analysis
Abstract
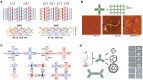
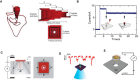
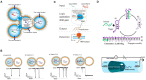
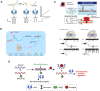
Nanopore technology is a promising label-free detection method. However, challenges exist for its further application in sequencing, clinical diagnostics and ultra-sensitive single molecule detection. The development of DNA nanotechnology nonetheless provides possible solutions to current obstacles hindering nanopore sensing technologies. In this review, we summarize recent relevant research contributing to efforts for developing nanopore methods associated with DNA nanotechnology. For example, DNA carriers can capture specific targets at pre-designed sites and escort them from nanopores at suitable speeds, thereby greatly enhancing capability and resolution for the detection of specific target molecules. In addition, DNA origami structures can be constructed to fulfill various design specifications and one-pot assembly reactions, thus serving as functional nanopores. Moreover, based on DNA strand displacement, nanopores can also be utilized to characterize the outputs of DNA computing and to develop programmable smart diagnostic nanodevices. In summary, DNA assembly-based nanopore research can pave the way for the realization of impactful biological detection and diagnostic platforms via single-biomolecule analysis.
© The Author(s) 2020. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures








References
-
- Feng J., Liu K., Bulushev R.D., Khlybov S., Dumcenco D., Kis A., Radenovic A.. Identification of single nucleotides in MoS2 nanopores. Nat. Nanotech. 2015; 10:1070–1076. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

