15N Stable Isotope Labeling and Comparative Metabolomics Facilitates Genome Mining in Cultured Cyanobacteria
- PMID: 32083834
- PMCID: PMC7297037
- DOI: 10.1021/acschembio.9b00993
15N Stable Isotope Labeling and Comparative Metabolomics Facilitates Genome Mining in Cultured Cyanobacteria
Abstract
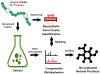
As genome mining becomes a more widely used approach to identify bacterial natural products, the challenge of matching biosynthetic gene clusters to their cognate secondary metabolites has become more apparent. Bioinformatic platforms such as AntiSMASH have made great progress in predicting chemical structures from genetic information, however the predicted structures are often incomplete. This complicates identifying the predicted compounds by mass spectrometry. Secondary metabolites produced by cyanobacteria represent a unique opportunity for bridging this gap. Cultured cyanobacteria incorporate inorganic nitrogen provided in chemically defined media into all nitrogen-containing secondary metabolites. Thus, stable isotope labeling with 15N labeled nitrate and subsequent comparative metabolomics can be used to match biosynthetic gene clusters to their cognate compounds in cell extracts. Analysis of the sequenced genome of Nostoc sp. UIC 10630 identified six biosynthetic gene clusters predicted to encode the production of a secondary metabolite with at least one nitrogen atom. Comparative metabolomic analysis of the 15N labeled and unlabeled cell extracts revealed four nitrogen containing compounds that contained the same number of nitrogen atoms as were predicted in the biosynthetic gene clusters. Two of the four compounds were new secondary metabolites, and their structures were elucidated by NMR, HRESIMS, and MS/MS.
Conflict of interest statement
The authors declare no competing financial interest.
Figures
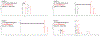


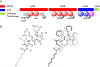
References
-
- Da Rocha AB, Lopes RM, and Schwartsmann G (2001) Natural products in anticancer therapy. Curr. Opin. Pharmacol 1, 364–369. - PubMed
-
- Newman DJ, and Cragg GM (2016) Natural Products as Sources of New Drugs from 1981 to 2014. J. Nat. Prod 79, 629–661. - PubMed
-
- Metzker ML (2010) Sequencing technologies the next generation. Nat. Rev. Genet 11, 31–46. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

