The fungal CCAAT-binding complex and HapX display highly variable but evolutionary conserved synergetic promoter-specific DNA recognition
- PMID: 32086516
- PMCID: PMC7144946
- DOI: 10.1093/nar/gkaa109
The fungal CCAAT-binding complex and HapX display highly variable but evolutionary conserved synergetic promoter-specific DNA recognition
Abstract
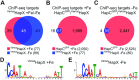
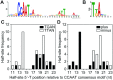
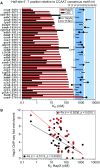
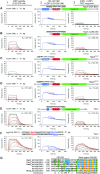
To sustain iron homeostasis, microorganisms have evolved fine-tuned mechanisms for uptake, storage and detoxification of the essential metal iron. In the human pathogen Aspergillus fumigatus, the fungal-specific bZIP-type transcription factor HapX coordinates adaption to both iron starvation and iron excess and is thereby crucial for virulence. Previous studies indicated that a HapX homodimer interacts with the CCAAT-binding complex (CBC) to cooperatively bind bipartite DNA motifs; however, the mode of HapX-DNA recognition had not been resolved. Here, combination of in vivo (genetics and ChIP-seq), in vitro (surface plasmon resonance) and phylogenetic analyses identified an astonishing plasticity of CBC:HapX:DNA interaction. DNA motifs recognized by the CBC:HapX protein complex comprise a bipartite DNA binding site 5'-CSAATN12RWT-3' and an additional 5'-TKAN-3' motif positioned 11-23 bp downstream of the CCAAT motif, i.e. occasionally overlapping the 3'-end of the bipartite binding site. Phylogenetic comparison taking advantage of 20 resolved Aspergillus species genomes revealed that DNA recognition by the CBC:HapX complex shows promoter-specific cross-species conservation rather than regulon-specific conservation. Moreover, we show that CBC:HapX interaction is absolutely required for all known functions of HapX. The plasticity of the CBC:HapX:DNA interaction permits fine tuning of CBC:HapX binding specificities that could support adaptation of pathogens to their host niches.
© The Author(s) 2020. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures





Similar articles
-
Deciphering the combinatorial DNA-binding code of the CCAAT-binding complex and the iron-regulatory basic region leucine zipper (bZIP) transcription factor HapX.J Biol Chem. 2015 Mar 6;290(10):6058-70. doi: 10.1074/jbc.M114.628677. Epub 2015 Jan 14. J Biol Chem. 2015. PMID: 25589790 Free PMC article.
-
The Janus transcription factor HapX controls fungal adaptation to both iron starvation and iron excess.EMBO J. 2014 Oct 1;33(19):2261-76. doi: 10.15252/embj.201489468. Epub 2014 Aug 4. EMBO J. 2014. PMID: 25092765 Free PMC article.
-
The bZIP Transcription Factor HapX Is Post-Translationally Regulated to Control Iron Homeostasis in Aspergillus fumigatus.Int J Mol Sci. 2021 Jul 20;22(14):7739. doi: 10.3390/ijms22147739. Int J Mol Sci. 2021. PMID: 34299357 Free PMC article.
-
The CCAAT-binding complex (CBC) in Aspergillus species.Biochim Biophys Acta Gene Regul Mech. 2017 May;1860(5):560-570. doi: 10.1016/j.bbagrm.2016.11.008. Epub 2016 Dec 8. Biochim Biophys Acta Gene Regul Mech. 2017. PMID: 27939757 Review.
-
Coordinated regulation of iron metabolism in Cryptococcus neoformans by GATA and CCAAT transcription factors: connections with virulence.Curr Genet. 2021 Aug;67(4):583-593. doi: 10.1007/s00294-021-01172-5. Epub 2021 Mar 24. Curr Genet. 2021. PMID: 33760942 Free PMC article. Review.
Cited by
-
HapX-mediated H2B deub1 and SreA-mediated H2A.Z deposition coordinate in fungal iron resistance.Nucleic Acids Res. 2023 Oct 27;51(19):10238-10260. doi: 10.1093/nar/gkad708. Nucleic Acids Res. 2023. PMID: 37650633 Free PMC article.
-
Yap5 Competes With Hap4 for the Regulation of Iron Homeostasis Genes in the Human Pathogen Candida glabrata.Front Cell Infect Microbiol. 2021 Nov 26;11:731988. doi: 10.3389/fcimb.2021.731988. eCollection 2021. Front Cell Infect Microbiol. 2021. PMID: 34900750 Free PMC article.
-
Exploring a novel genomic safe-haven site in the human pathogenic mould Aspergillus fumigatus.Fungal Genet Biol. 2022 Jul;161:103702. doi: 10.1016/j.fgb.2022.103702. Epub 2022 May 13. Fungal Genet Biol. 2022. PMID: 35569804 Free PMC article.
-
First Investigative Study of Azole-Resistant Aspergillus fumigatus in the Environment in Burkina Faso.Int J Environ Res Public Health. 2021 Feb 25;18(5):2250. doi: 10.3390/ijerph18052250. Int J Environ Res Public Health. 2021. PMID: 33668719 Free PMC article.
-
Regulation of Ergosterol Biosynthesis in Pathogenic Fungi: Opportunities for Therapeutic Development.Microorganisms. 2025 Apr 10;13(4):862. doi: 10.3390/microorganisms13040862. Microorganisms. 2025. PMID: 40284698 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

