Early phylogenetic estimate of the effective reproduction number of SARS-CoV-2
- PMID: 32096566
- PMCID: PMC7228357
- DOI: 10.1002/jmv.25723
Early phylogenetic estimate of the effective reproduction number of SARS-CoV-2
Abstract
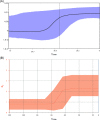
To reconstruct the evolutionary dynamics of the 2019 novel-coronavirus recently causing an outbreak in Wuhan, China, 52 SARS-CoV-2 genomes available on 4 February 2020 at Global Initiative on Sharing All Influenza Data were analyzed. The two models used to estimate the reproduction number (coalescent-based exponential growth and a birth-death skyline method) indicated an estimated mean evolutionary rate of 7.8 × 10-4 subs/site/year (range, 1.1 × 10-4 -15 × 10-4 ) and a mean tMRCA of the tree root of 73 days. The estimated R value was 2.6 (range, 2.1-5.1), and increased from 0.8 to 2.4 in December 2019. The estimated mean doubling time of the epidemic was between 3.6 and 4.1 days. This study proves the usefulness of phylogeny in supporting the surveillance of emerging new infections even as the epidemic is growing.
Keywords: SARS-CoV-2; evolutionary dynamics; reproductive number.
© 2020 Wiley Periodicals, Inc.
Conflict of interest statement
The authors declare that there are no conflict of interests.
Figures
References
-
- Posada D. jModelTest: phylogenetic model averaging. Mol Biol Evol. 2008;25(7):1253‐1256. - PubMed

