Parainfluenza virus 5 fusion protein maintains pre-fusion stability but not fusogenic activity following mutation of a transmembrane leucine/isoleucine domain
- PMID: 32100701
- PMCID: PMC7414451
- DOI: 10.1099/jgv.0.001399
Parainfluenza virus 5 fusion protein maintains pre-fusion stability but not fusogenic activity following mutation of a transmembrane leucine/isoleucine domain
Abstract
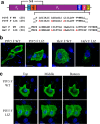
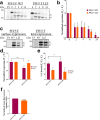
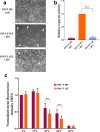
The paramyxoviruses Hendra virus (HeV) and parainfluenza virus 5 (PIV5) require the fusion (F) protein to efficiently infect cells. For fusion to occur, F undergoes dramatic, essentially irreversible conformational changes to merge the viral and cell membranes into a continuous bilayer. Recently, a transmembrane (TM) domain leucine/isoleucine (L/I) zipper was shown to be critical in maintaining the expression, stability and pre-fusion conformation of HeV F, allowing for fine-tuned timing of membrane fusion. To analyse the effect of the TM domain L/I zipper in another paramyxovirus, we created alanine mutations to the TM domain of PIV5 F, a paramyxovirus model system. Our data show that while the PIV5 F TM L/I zipper does not significantly affect total expression and only modestly affects surface expression and pre-fusion stability, it is critical for fusogenic activity. These results suggest that the roles of TM L/I zipper motifs differ among members of the family Paramyxoviridae.
Keywords: fusogenic activity; leucine/isoleucine zipper; pre-fusion stability; transmembrane domain.
Conflict of interest statement
The authors declare that there are no conflicts of interest.
Figures
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

