Comparison of Shiga toxin-encoding bacteriophages in highly pathogenic strains of Shiga toxin-producing Escherichia coli O157:H7 in the UK
- PMID: 32100710
- PMCID: PMC7200060
- DOI: 10.1099/mgen.0.000334
Comparison of Shiga toxin-encoding bacteriophages in highly pathogenic strains of Shiga toxin-producing Escherichia coli O157:H7 in the UK
Abstract
Over the last 35 years in the UK, the burden of Shiga toxin-producing Escherichia coli (STEC) O157:H7 infection has, during different periods of time, been associated with five different sub-lineages (1983-1995, Ia, I/IIa and I/IIb; 1996-2014, Ic; and 2015-2018, IIb). The acquisition of a stx2a-encoding bacteriophage by these five sub-lineages appears to have coincided with their respective emergences. The Oxford Nanopore Technologies (ONT) system was used to sequence, characterize and compare the stx-encoding prophages harboured by each sub-lineage to investigate the integration of this key virulence factor. The stx2a-encoding prophages from each of the lineages causing clinical disease in the UK were all different, including the two UK sub-lineages (Ia and I/IIa) circulating concurrently and causing severe disease in the early 1980s. Comparisons between the stx2a-encoding prophage in sub-lineages I/IIb and IIb revealed similarity to the prophage commonly found to encode stx2c, and the same site of bacteriophage integration (sbcB) as stx2c-encoding prophage. These data suggest independent acquisition of previously unobserved stx2a-encoding phage is more likely to have contributed to the emergence of STEC O157:H7 sub-lineages in the UK than intra-UK lineage to lineage phage transmission. In contrast, the stx2c-encoding prophage showed a high level of similarity across lineages and time, consistent with the model of stx2c being present in the common ancestor to extant STEC O157:H7 and maintained by vertical inheritance in the majority of the population. Studying the nature of the stx-encoding bacteriophage contributes to our understanding of the emergence of highly pathogenic strains of STEC O157:H7.
Keywords: Escherichia coli O157:H7; Shiga toxin; bacteriophage; whole-genome sequencing.
Conflict of interest statement
The authors declare that there are no conflicts of interest.
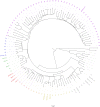
Figures





Similar articles
-
Shiga Toxin (Stx) Phage-Encoded Lytic Genes Are Not Required for the Mouse Virulence of O157:H7 Escherichia coli Stx2-Producing Clinical Isolates.Microbiol Spectr. 2023 Jun 15;11(3):e0037223. doi: 10.1128/spectrum.00372-23. Epub 2023 Apr 6. Microbiol Spectr. 2023. PMID: 37022201 Free PMC article.
-
Escherichia coli O157:H7 strains harbor at least three distinct sequence types of Shiga toxin 2a-converting phages.BMC Genomics. 2015 Sep 29;16:733. doi: 10.1186/s12864-015-1934-1. BMC Genomics. 2015. PMID: 26416807 Free PMC article.
-
Re-analysis of an outbreak of Shiga toxin-producing Escherichia coli O157:H7 associated with raw drinking milk using Nanopore sequencing.Sci Rep. 2024 Mar 9;14(1):5821. doi: 10.1038/s41598-024-54662-0. Sci Rep. 2024. PMID: 38461188 Free PMC article.
-
Phage applications for biocontrol of enterohemorrhagic E. coli O157:H7 and other Shiga toxin-producing Escherichia coli.Int J Food Microbiol. 2025 Aug 2;439:111267. doi: 10.1016/j.ijfoodmicro.2025.111267. Epub 2025 May 13. Int J Food Microbiol. 2025. PMID: 40382813 Review.
-
A Toxic Environment: a Growing Understanding of How Microbial Communities Affect Escherichia coli O157:H7 Shiga Toxin Expression.Appl Environ Microbiol. 2020 Nov 24;86(24):e00509-20. doi: 10.1128/AEM.00509-20. Print 2020 Nov 24. Appl Environ Microbiol. 2020. PMID: 32358004 Free PMC article. Review.
Cited by
-
Characterization of a pESI-like plasmid and analysis of multidrug-resistant Salmonella enterica Infantis isolates in England and Wales.Microb Genom. 2021 Oct;7(10):000658. doi: 10.1099/mgen.0.000658. Microb Genom. 2021. PMID: 34647862 Free PMC article.
-
Characterization of a P1-bacteriophage-like plasmid (phage-plasmid) harbouring bla CTX-M-15 in Salmonella enterica serovar Typhi.Microb Genom. 2022 Dec;8(12):mgen000913. doi: 10.1099/mgen.0.000913. Microb Genom. 2022. PMID: 36748517 Free PMC article.
-
Comparative Genome Analysis Reveals Accumulation of Single-Nucleotide Repeats in Pathogenic Escherichia Lineages.Curr Issues Mol Biol. 2022 Jan 20;44(2):498-504. doi: 10.3390/cimb44020034. Curr Issues Mol Biol. 2022. PMID: 35723320 Free PMC article.
-
Comparative Genomics of Escherichia coli Serotype O55:H7 Using Complete Closed Genomes.Microorganisms. 2022 Jul 30;10(8):1545. doi: 10.3390/microorganisms10081545. Microorganisms. 2022. PMID: 36013963 Free PMC article.
-
Bacteriophages of Shiga Toxin-Producing Escherichia coli and Their Contribution to Pathogenicity.Pathogens. 2021 Mar 29;10(4):404. doi: 10.3390/pathogens10040404. Pathogens. 2021. PMID: 33805526 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

