PDGFR-β Promoter Forms a Vacancy G-Quadruplex that Can Be Filled in by dGMP: Solution Structure and Molecular Recognition of Guanine Metabolites and Drugs
- PMID: 32101424
- PMCID: PMC7241120
- DOI: 10.1021/jacs.9b12770
PDGFR-β Promoter Forms a Vacancy G-Quadruplex that Can Be Filled in by dGMP: Solution Structure and Molecular Recognition of Guanine Metabolites and Drugs
Abstract
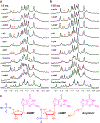
Aberrant expression of PDGFR-β is associated with a number of diseases. The G-quadruplexes (G4s) formed in PDGFR-β gene promoter are transcriptional modulators and amenable to small molecule targeting. The major G4 formed in the PDGFR-β gene promoter was previously shown to have a broken G-strand. Herein, we report that the PDGFR-β gene promoter sequence forms a vacancy G-quadruplex (vG4) which can be filled in and stabilized by physiologically relevant guanine metabolites, such as dGMP, GMP, and cGMP, as well as guanine-derivative drugs. We determined the NMR structure of the dGMP-fill-in PDGFR-β vG4 in K+ solution. This is the first structure of a guanine-metabolite-fill-in vG4 based on a human gene promoter sequence. Our structure and systematic analysis elucidate the contributions of Hoogsten hydrogen bonds, sugar, and phosphate moieties to the specific G-vacancy fill-in. Intriguingly, an equilibrium of 3'- and 5'-end vG4s is present in the PDGFR-β promoter sequence, and dGMP favors the 5'-end fill-in. Guanine metabolites and drugs were tested and showed a conserved selectivity for the 5'-vacancy, except for cGMP. cGMP binds both the 3'- and 5'-end vG4s and forms two fill-in G4s with similar population. Significantly, guanine metabolites are involved in many physiological and pathological processes in human cells; thus, our results provide a structural basis to understand their potential regulatory functions by interaction with promoter vG4s. Moreover, the NMR structure can guide rational design of ligands that target the PDGFR-β vG4.
Conflict of interest statement
The authors declare no competing interests.
Figures


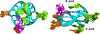

Similar articles
-
Solution Structure of Ternary Complex of Berberine Bound to a dGMP-Fill-In Vacancy G-Quadruplex Formed in the PDGFR-β Promoter.J Am Chem Soc. 2021 Oct 13;143(40):16549-16555. doi: 10.1021/jacs.1c06200. Epub 2021 Sep 29. J Am Chem Soc. 2021. PMID: 34586799 Free PMC article.
-
Oxidative Damage Induces a Vacancy G-Quadruplex That Binds Guanine Metabolites: Solution Structure of a cGMP Fill-in Vacancy G-Quadruplex in the Oxidized BLM Gene Promoter.J Am Chem Soc. 2022 Apr 13;144(14):6361-6372. doi: 10.1021/jacs.2c00435. Epub 2022 Mar 30. J Am Chem Soc. 2022. PMID: 35352895 Free PMC article.
-
DNA G-Quadruplex in Human Telomeres and Oncogene Promoters: Structures, Functions, and Small Molecule Targeting.Acc Chem Res. 2022 Sep 20;55(18):2628-2646. doi: 10.1021/acs.accounts.2c00337. Epub 2022 Sep 2. Acc Chem Res. 2022. PMID: 36054116 Free PMC article.
-
Specific Recognition of Promoter G-Quadruplex DNAs by Small Molecule Ligands and Light-up Probes.ACS Chem Biol. 2019 Oct 18;14(10):2102-2114. doi: 10.1021/acschembio.9b00475. Epub 2019 Oct 4. ACS Chem Biol. 2019. PMID: 31532996 Review.
-
Structure of the biologically relevant G-quadruplex in the c-MYC promoter.Nucleosides Nucleotides Nucleic Acids. 2006;25(8):951-68. doi: 10.1080/15257770600809913. Nucleosides Nucleotides Nucleic Acids. 2006. PMID: 16901825 Review.
Cited by
-
Quadruplex-Duplex Junction: A High-Affinity Binding Site for Indoloquinoline Ligands.Chemistry. 2020 Dec 15;26(70):16910-16922. doi: 10.1002/chem.202003540. Epub 2020 Nov 16. Chemistry. 2020. PMID: 32975874 Free PMC article.
-
G-quadruplexes on chromosomal DNA negatively regulates topoisomerase 1 activity.Nucleic Acids Res. 2024 Mar 21;52(5):2142-2156. doi: 10.1093/nar/gkae073. Nucleic Acids Res. 2024. PMID: 38340342 Free PMC article.
-
Custom G4 Microarrays Reveal Selective G-Quadruplex Recognition of Small Molecule BMVC: A Large-Scale Assessment of Ligand Binding Selectivity.Molecules. 2020 Jul 30;25(15):3465. doi: 10.3390/molecules25153465. Molecules. 2020. PMID: 32751510 Free PMC article.
-
Solution Structure of Ternary Complex of Berberine Bound to a dGMP-Fill-In Vacancy G-Quadruplex Formed in the PDGFR-β Promoter.J Am Chem Soc. 2021 Oct 13;143(40):16549-16555. doi: 10.1021/jacs.1c06200. Epub 2021 Sep 29. J Am Chem Soc. 2021. PMID: 34586799 Free PMC article.
-
Structural basis for high-affinity recognition of aflatoxin B1 by a DNA aptamer.Nucleic Acids Res. 2023 Aug 11;51(14):7666-7674. doi: 10.1093/nar/gkad541. Nucleic Acids Res. 2023. PMID: 37351632 Free PMC article.
References
-
- Hänsel-Hertsch R; Beraldi D; Lensing SV; Marsico G; Zyner K; Parry A; Di Antonio M; Pike J; Kimura H; Narita M, G-quadruplex structures mark human regulatory chromatin. Nat. Genet 2016, 48 (10), 1267–1275. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

