A neural network for online spike classification that improves decoding accuracy
- PMID: 32101491
- PMCID: PMC7191521
- DOI: 10.1152/jn.00641.2019
A neural network for online spike classification that improves decoding accuracy
Abstract
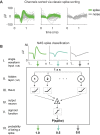
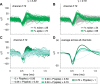
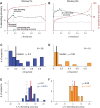
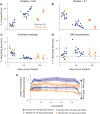
Separating neural signals from noise can improve brain-computer interface performance and stability. However, most algorithms for separating neural action potentials from noise are not suitable for use in real time and have shown mixed effects on decoding performance. With the goal of removing noise that impedes online decoding, we sought to automate the intuition of human spike-sorters to operate in real time with an easily tunable parameter governing the stringency with which spike waveforms are classified. We trained an artificial neural network with one hidden layer on neural waveforms that were hand-labeled as either spikes or noise. The network output was a likelihood metric for each waveform it classified, and we tuned the network's stringency by varying the minimum likelihood value for a waveform to be considered a spike. Using the network's labels to exclude noise waveforms, we decoded remembered target location during a memory-guided saccade task from electrode arrays implanted in prefrontal cortex of rhesus macaque monkeys. The network classified waveforms in real time, and its classifications were qualitatively similar to those of a human spike-sorter. Compared with decoding with threshold crossings, in most sessions we improved decoding performance by removing waveforms with low spike likelihood values. Furthermore, decoding with our network's classifications became more beneficial as time since array implantation increased. Our classifier serves as a feasible preprocessing step, with little risk of harm, that could be applied to both off-line neural data analyses and online decoding.NEW & NOTEWORTHY Although there are many spike-sorting methods that isolate well-defined single units, these methods typically involve human intervention and have inconsistent effects on decoding. We used human classified neural waveforms as training data to create an artificial neural network that could be tuned to separate spikes from noise that impaired decoding. We found that this network operated in real time and was suitable for both off-line data processing and online decoding.
Keywords: BCI; decoding; neural network; prefrontal cortex; spike-sorting.
Conflict of interest statement
No conflicts of interest, financial or otherwise, are declared by the authors.
Figures

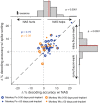
References
-
- Abadi M, Agarwal A, Barham P, Brevdo E, Chen Z, Citro C, Corrado GS, Davis A, Dean J, Devin M, Ghemawat S, Goodfellow I, Harp A, Irving G, Isard M, Jia Y, Jozefowicz R, Kaiser L, Kudlur M, Levenberg J, Mane D, Monga R, Moore S, Murray D, Olah C, Schuster M, Shlens J, Steiner B, Sutskever I, Talwar K, Tucker P, Vanhoucke V, Vasudevan V, Viegas F, Vinyals O, Warden P, Wattenberg M, Wicke M, Yu Y, Zheng X. TensorFlow: large-scale machine learning on heterogeneous distributed systems. In: OSDI'16: Proceedings of the 12th USENIX Conference on Operating Systems ... Berkeley, CA: USENIX Association, 2016.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

