A Systematic Approach to Bacterial Phylogeny Using Order Level Sampling and Identification of HGT Using Network Science
- PMID: 32102454
- PMCID: PMC7074868
- DOI: 10.3390/microorganisms8020312
A Systematic Approach to Bacterial Phylogeny Using Order Level Sampling and Identification of HGT Using Network Science
Abstract
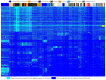
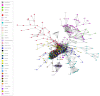
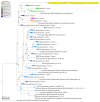
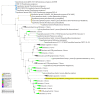
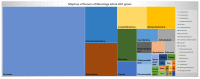
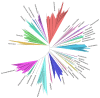
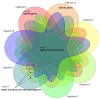
Reconstructing and visualizing phylogenetic relationships among living organisms is a fundamental challenge because not all organisms share the same genes. As a result, the first phylogenetic visualizations employed a single gene, e.g., rRNA genes, sufficiently conserved to be present in all organisms but divergent enough to provide discrimination between groups. As more genome data became available, researchers began concatenating different combinations of genes or proteins to construct phylogenetic trees believed to be more robust because they incorporated more information. However, the genes or proteins chosen were based on ad hoc approaches. The large number of complete genome sequences available today allows the use of whole genomes to analyze relationships among organisms rather than using an ad hoc set of genes. We present a systematic approach for constructing a phylogenetic tree based on simultaneously clustering the complete proteomes of 360 bacterial species. From the homologous clusters, we identify 49 protein sequences shared by 99% of the organisms to build a tree. Of the 49 sequences, 47 have homologous sequences in both archaea and eukarya. The clusters are also used to create a network from which bacterial species with horizontally-transferred genes from other phyla are identified.
Keywords: horizontal gene transfer; network of bacteria; network science; phylogeny; tree of bacterial phyla.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
Similar articles
-
Genome trees constructed using five different approaches suggest new major bacterial clades.BMC Evol Biol. 2001 Oct 20;1:8. doi: 10.1186/1471-2148-1-8. BMC Evol Biol. 2001. PMID: 11734060 Free PMC article.
-
The effects of model choice and mitigating bias on the ribosomal tree of life.Mol Phylogenet Evol. 2013 Oct;69(1):17-38. doi: 10.1016/j.ympev.2013.05.006. Epub 2013 May 22. Mol Phylogenet Evol. 2013. PMID: 23707703
-
A database of phylogenetically atypical genes in archaeal and bacterial genomes, identified using the DarkHorse algorithm.BMC Bioinformatics. 2008 Oct 7;9:419. doi: 10.1186/1471-2105-9-419. BMC Bioinformatics. 2008. PMID: 18840280 Free PMC article.
-
The tree of life viewed through the contents of genomes.Methods Mol Biol. 2009;532:141-61. doi: 10.1007/978-1-60327-853-9_8. Methods Mol Biol. 2009. PMID: 19271183 Review.
-
Proteogenomics of rare taxonomic phyla: A prospective treasure trove of protein coding genes.Proteomics. 2016 Jan;16(2):226-40. doi: 10.1002/pmic.201500263. Epub 2015 Nov 23. Proteomics. 2016. PMID: 26773550 Review.
Cited by
-
Frequent transitions in self-assembly across the evolution of a central metabolic enzyme.bioRxiv [Preprint]. 2024 Jul 7:2024.07.05.602260. doi: 10.1101/2024.07.05.602260. bioRxiv. 2024. Update in: Nat Commun. 2024 Dec 3;15(1):10515. doi: 10.1038/s41467-024-54408-6. PMID: 39005358 Free PMC article. Updated. Preprint.
-
Sequence determinants of human-cell entry identified in ACE2-independent bat sarbecoviruses: A combined laboratory and computational network science approach.EBioMedicine. 2022 May;79:103990. doi: 10.1016/j.ebiom.2022.103990. Epub 2022 Apr 8. EBioMedicine. 2022. PMID: 35405384 Free PMC article.
-
Frequent transitions in self-assembly across the evolution of a central metabolic enzyme.Nat Commun. 2024 Dec 3;15(1):10515. doi: 10.1038/s41467-024-54408-6. Nat Commun. 2024. PMID: 39627196 Free PMC article.
-
Obligate movements of an active site-linked surface domain control RNA polymerase elongation and pausing via a Phe pocket anchor.Proc Natl Acad Sci U S A. 2021 Sep 7;118(36):e2101805118. doi: 10.1073/pnas.2101805118. Proc Natl Acad Sci U S A. 2021. PMID: 34470825 Free PMC article.
-
Respirable Metals, Bacteria, and Fungi during a Saharan-Sahelian Dust Event in Houston, Texas.Environ Sci Technol. 2023 Dec 5;57(48):19942-19955. doi: 10.1021/acs.est.3c04158. Epub 2023 Nov 9. Environ Sci Technol. 2023. PMID: 37943153 Free PMC article.
References
-
- Easley D., Kleinberg J. Networks, Crowds, and Markets. Volume 8 Cambridge University Press; Cambridge, UK: 2010.
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

