The luminescent HiBiT peptide enables selective quantitation of G protein-coupled receptor ligand engagement and internalization in living cells
- PMID: 32107310
- PMCID: PMC7152755
- DOI: 10.1074/jbc.RA119.011952
The luminescent HiBiT peptide enables selective quantitation of G protein-coupled receptor ligand engagement and internalization in living cells
Abstract
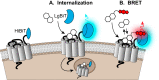
G protein-coupled receptors (GPCRs) are prominent targets to new therapeutics for a range of diseases. Comprehensive assessments of their cellular interactions with bioactive compounds, particularly in a kinetic format, are imperative to the development of drugs with improved efficacy. Hence, we developed complementary cellular assays that enable equilibrium and real-time analyses of GPCR ligand engagement and consequent activation, measured as receptor internalization. These assays utilize GPCRs genetically fused to an N-terminal HiBiT peptide (1.3 kDa), which produces bright luminescence upon high-affinity complementation with LgBiT, an 18-kDa subunit derived from NanoLuc. The cell impermeability of LgBiT limits signal detection to the cell surface and enables measurements of ligand-induced internalization through changes in cell-surface receptor density. In addition, bioluminescent resonance energy transfer is used to quantify dynamic interactions between ligands and their cognate HiBiT-tagged GPCRs through competitive binding with fluorescent tracers. The sensitivity and dynamic range of these assays benefit from the specificity of bioluminescent resonance energy transfer and the high signal intensity of HiBiT/LgBiT without background luminescence from receptors present in intracellular compartments. These features allow analyses of challenging interactions having low selectivity or affinity and enable studies using endogenously tagged receptors. Using the β-adrenergic receptor family as a model, we demonstrate the versatility of these assays by utilizing the same HiBiT construct in analyses of multiple aspects of GPCR pharmacology. We anticipate that this combination of target engagement and proximal functional readout will prove useful to the study of other GPCR families and the development of new therapeutics.
Keywords: G protein-coupled receptor (GPCR); HiBiT; NanoLuc; adrenergic receptor; bioluminescence; bioluminescence resonance energy transfer (BRET); endogenous tagging; kinetics; ligand engagement; receptor internalization.
© 2020 Boursier et al.
Conflict of interest statement
All authors were Promega employees during manuscript preparation
Figures




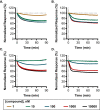
References
-
- Schuetz D. A., de Witte W. E. A., Wong Y. C., Knasmueller B., Richter L., Kokh D. B., Sadiq S. K., Bosma R., Nederpelt I., Heitman L. H., Segala E., Amaral M., Guo D., Andres D., Georgi V., et al. (2017) Kinetics for drug discovery: an industry driven effort to target drug residence time. Drug Discov. Today 22, 896–911 10.1016/j.drudis.2017.02.002 - DOI - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

