Control of tissue morphogenesis by the HOX gene Ultrabithorax
- PMID: 32122911
- PMCID: PMC7063672
- DOI: 10.1242/dev.184564
Control of tissue morphogenesis by the HOX gene Ultrabithorax
Abstract
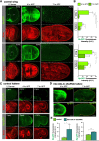
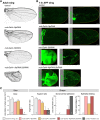
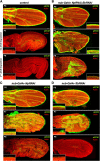
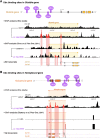
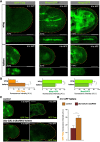
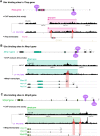
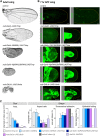
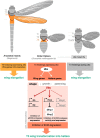
Mutations in the Ultrabithorax (Ubx) gene cause homeotic transformation of the normally two-winged Drosophila into a four-winged mutant fly. Ubx encodes a HOX family transcription factor that specifies segment identity, including transformation of the second set of wings into rudimentary halteres. Ubx is known to control the expression of many genes that regulate tissue growth and patterning, but how it regulates tissue morphogenesis to reshape the wing into a haltere is still unclear. Here, we show that Ubx acts by repressing the expression of two genes in the haltere, Stubble and Notopleural, both of which encode transmembrane proteases that remodel the apical extracellular matrix to promote wing morphogenesis. In addition, Ubx induces expression of the Tissue inhibitor of metalloproteases in the haltere, which prevents the basal extracellular matrix remodelling necessary for wing morphogenesis. Our results provide a long-awaited explanation for how Ubx controls morphogenetic transformation.
Keywords: Development; Drosophila; Morphogenesis; Notopleural; Stubble.
© 2020. Published by The Company of Biologists Ltd.
Conflict of interest statement
Competing interestsThe authors declare no competing or financial interests.
Figures
Comment in
-
Evo-Devo: When Four Became Two Plus Two.Curr Biol. 2020 Jun 8;30(11):R655-R657. doi: 10.1016/j.cub.2020.04.022. Curr Biol. 2020. PMID: 32516617
Similar articles
-
Hox gene Ultrabithorax regulates distinct sets of target genes at successive stages of Drosophila haltere morphogenesis.Proc Natl Acad Sci U S A. 2011 Feb 15;108(7):2855-60. doi: 10.1073/pnas.1015077108. Epub 2011 Jan 31. Proc Natl Acad Sci U S A. 2011. PMID: 21282633 Free PMC article.
-
Negative regulation of dorsoventral signaling by the homeotic gene Ultrabithorax during haltere development in Drosophila.Dev Biol. 1999 Aug 15;212(2):491-502. doi: 10.1006/dbio.1999.9341. Dev Biol. 1999. PMID: 10433837
-
The Drosophila Hox gene Ultrabithorax controls appendage shape by regulating extracellular matrix dynamics.Development. 2018 Jul 9;145(13):dev161844. doi: 10.1242/dev.161844. Development. 2018. PMID: 29853618
-
Ultrabithorax and the evolution of insect forewing/hindwing differentiation.Curr Opin Insect Sci. 2017 Feb;19:8-15. doi: 10.1016/j.cois.2016.10.007. Epub 2016 Nov 2. Curr Opin Insect Sci. 2017. PMID: 28521947 Review.
-
Haltere development in D. melanogaster: implications for the evolution of appendage size, shape and function.Int J Dev Biol. 2020;64(1-2-3):159-165. doi: 10.1387/ijdb.190133LS. Int J Dev Biol. 2020. PMID: 32659004 Review.
Cited by
-
Regulatory functions of homeobox domain transcription factors in fungi.Appl Environ Microbiol. 2024 Mar 20;90(3):e0220823. doi: 10.1128/aem.02208-23. Epub 2024 Feb 29. Appl Environ Microbiol. 2024. PMID: 38421174 Free PMC article. Review.
-
Spatiotemporal remodeling of extracellular matrix orients epithelial sheet folding.Sci Adv. 2023 Sep;9(35):eadh2154. doi: 10.1126/sciadv.adh2154. Epub 2023 Sep 1. Sci Adv. 2023. PMID: 37656799 Free PMC article.
-
Role of the Hox Genes, Sex combs reduced, Fushi tarazu and Antennapedia, in Leg Development of the Spider Mite Tetranychus urticae.Int J Mol Sci. 2023 Jun 20;24(12):10391. doi: 10.3390/ijms241210391. Int J Mol Sci. 2023. PMID: 37373537 Free PMC article.
-
Characterisation of the role and regulation of Ultrabithorax in sculpting fine-scale leg morphology.Front Cell Dev Biol. 2023 Feb 13;11:1119221. doi: 10.3389/fcell.2023.1119221. eCollection 2023. Front Cell Dev Biol. 2023. PMID: 36861038 Free PMC article.
-
Form and function of the apical extracellular matrix: new insights from Caenorhabditis elegans, Drosophila melanogaster, and the vertebrate inner ear.Fac Rev. 2020 Dec 22;9:27. doi: 10.12703/r/9-27. eCollection 2020. Fac Rev. 2020. PMID: 33659959 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

