Sequential Glycosylation of Proteins with Substrate-Specific N-Glycosyltransferases
- PMID: 32123732
- PMCID: PMC7047269
- DOI: 10.1021/acscentsci.9b00021
Sequential Glycosylation of Proteins with Substrate-Specific N-Glycosyltransferases
Abstract
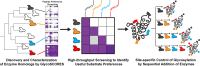
Protein glycosylation is a common post-translational modification that influences the functions and properties of proteins. Despite advances in methods to produce defined glycoproteins by chemoenzymatic elaboration of monosaccharides, the understanding and engineering of glycoproteins remain challenging, in part, due to the difficulty of site-specifically controlling glycosylation at each of several positions within a protein. Here, we address this limitation by discovering and exploiting the unique, conditionally orthogonal peptide acceptor specificities of N-glycosyltransferases (NGTs). We used cell-free protein synthesis and mass spectrometry of self-assembled monolayers to rapidly screen 41 putative NGTs and rigorously characterize the unique acceptor sequence preferences of four NGT variants using 1254 acceptor peptides and 8306 reaction conditions. We then used the optimized NGT-acceptor sequence pairs to sequentially install monosaccharides at four sites within one target protein. This strategy to site-specifically control the installation of N-linked monosaccharides for elaboration to a variety of functional N-glycans overcomes a major limitation in synthesizing defined glycoproteins for research and therapeutic applications.
Copyright © 2020 American Chemical Society.
Conflict of interest statement
The authors declare no competing financial interest.
Figures




References
-
- Elliott S.; Lorenzini T.; Asher S.; Aoki K.; Brankow D.; Buck L.; Busse L.; Chang D.; Fuller J.; Grant J.; Hernday N.; Hokum M.; Hu S.; Knudten A.; Levin N.; Komorowski R.; Martin F.; Navarro R.; Osslund T.; Rogers G.; Rogers N.; Trail G.; Egrie J. Enhancement of therapeutic protein in vivo activities through glycoengineering. Nat. Biotechnol. 2003, 21 (4), 414–421. 10.1038/nbt799. - DOI - PubMed
-
- Lin C.-W.; Tsai M.-H.; Li S.-T.; Tsai T.-I.; Chu K.-C.; Liu Y.-C.; Lai M.-Y.; Wu C.-Y.; Tseng Y.-C.; Shivatare S. S.; Wang C.-H.; Chao P.; Wang S.-Y.; Shih H.-W.; Zeng Y.-F.; You T.-H.; Liao J.-Y.; Tu Y.-C.; Lin Y.-S.; Chuang H.-Y.; Chen C.-L.; Tsai C.-S.; Huang C.-C.; Lin N.-H.; Ma C.; Wu C.-Y.; Wong C.-H. A common glycan structure on immunoglobulin g for enhancement of effector functions. Proc. Natl. Acad. Sci. U. S. A. 2015, 112 (34), 10611–10616. 10.1073/pnas.1513456112. - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources

