Transcriptomic evidence that von Economo neurons are regionally specialized extratelencephalic-projecting excitatory neurons
- PMID: 32127543
- PMCID: PMC7054400
- DOI: 10.1038/s41467-020-14952-3
Transcriptomic evidence that von Economo neurons are regionally specialized extratelencephalic-projecting excitatory neurons
Abstract
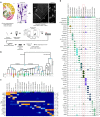
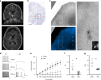
von Economo neurons (VENs) are bipolar, spindle-shaped neurons restricted to layer 5 of human frontoinsula and anterior cingulate cortex that appear to be selectively vulnerable to neuropsychiatric and neurodegenerative diseases, although little is known about other VEN cellular phenotypes. Single nucleus RNA-sequencing of frontoinsula layer 5 identifies a transcriptomically-defined cell cluster that contained VENs, but also fork cells and a subset of pyramidal neurons. Cross-species alignment of this cell cluster with a well-annotated mouse classification shows strong homology to extratelencephalic (ET) excitatory neurons that project to subcerebral targets. This cluster also shows strong homology to a putative ET cluster in human temporal cortex, but with a strikingly specific regional signature. Together these results suggest that VENs are a regionally distinctive type of ET neuron. Additionally, we describe the first patch clamp recordings of VENs from neurosurgically-resected tissue that show distinctive intrinsic membrane properties relative to neighboring pyramidal neurons.
Conflict of interest statement
The authors declare no competing interests.
Figures



References
-
- Von Economo C. A new type of special cells of the cingulate and insular lobes. Z. Ges. Neurol. Psychiatr. 1926;100:706–712. doi: 10.1007/BF02970950. - DOI
-
- Raghanti, M. A. et al. A comparison of the cortical structure of the bowhead whale (Balaena mysticetus), a Basal Mysticete, with Other Cetaceans. Anat. Rec. (Hoboken) 302, 745–760 (2018). - PubMed

