Characterization of the Human KCNQ1 Voltage Sensing Domain (VSD) in Lipodisq Nanoparticles for Electron Paramagnetic Resonance (EPR) Spectroscopic Studies of Membrane Proteins
- PMID: 32130007
- PMCID: PMC7202405
- DOI: 10.1021/acs.jpcb.9b11506
Characterization of the Human KCNQ1 Voltage Sensing Domain (VSD) in Lipodisq Nanoparticles for Electron Paramagnetic Resonance (EPR) Spectroscopic Studies of Membrane Proteins
Abstract
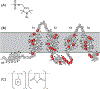
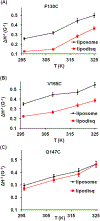
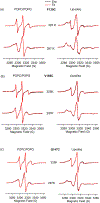
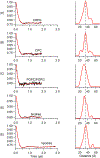
Membrane proteins are responsible for conducting essential biological functions that are necessary for the survival of living organisms. In spite of their physiological importance, limited structural information is currently available as a result of challenges in applying biophysical techniques for studying these protein systems. Electron paramagnetic resonance (EPR) spectroscopy is a very powerful technique to study the structural and dynamic properties of membrane proteins. However, the application of EPR spectroscopy to membrane proteins in a native membrane-bound state is extremely challenging due to the complexity observed in inhomogeneity sample preparation and the dynamic motion of the spin label. Detergent micelles are very popular membrane mimetics for membrane proteins due to their smaller size and homogeneity, providing high-resolution structure analysis by solution NMR spectroscopy. However, it is important to test whether the protein structure in a micelle environment is the same as that of its membrane-bound state. Lipodisq nanoparticles or styrene-maleic acid copolymer-lipid nanoparticles (SMALPs) have been introduced as a potentially good membrane-mimetic system for structural studies of membrane proteins. Recently, we reported on the EPR characterization of the KCNE1 membrane protein having a single transmembrane incorporated into lipodisq nanoparticles. In this work, lipodisq nanoparticles were used as a membrane mimic system for probing the structural and dynamic properties of the more complicated membrane protein system human KCNQ1 voltage sensing domain (Q1-VSD) having four transmembrane helices using site-directed spin-labeling EPR spectroscopy. Characterization of spin-labeled Q1-VSD incorporated into lipodisq nanoparticles was carried out using CW-EPR spectral line shape analysis and pulsed EPR double-electron electron resonance (DEER) measurements. The CW-EPR spectra indicate an increase in spectral line broadening with the addition of the styrene-maleic acid (SMA) polymer which approaches close to the rigid limit providing a homogeneous stabilization of the protein-lipid complex. Similarly, EPR DEER measurements indicated a superior quality of distance measurement with an increase in the phase memory time (Tm) values upon incorporation of the sample into lipodisq nanoparticles when compared to proteoliposomes. These results are consistent with the solution NMR structural studies on the Q1-VSD. This study will be beneficial for researchers working on investigating the structural and dynamic properties of more complicated membrane protein systems using lipodisq nanoparticles.
Conflict of interest statement
The authors declare no competing financial interest.
Figures

Similar articles
-
Electron paramagnetic resonance spectroscopic characterization of the human KCNE3 protein in lipodisq nanoparticles for structural dynamics of membrane proteins.Biophys Chem. 2023 Oct;301:107080. doi: 10.1016/j.bpc.2023.107080. Epub 2023 Jul 26. Biophys Chem. 2023. PMID: 37531799 Free PMC article.
-
Characterization of KCNE1 inside Lipodisq Nanoparticles for EPR Spectroscopic Studies of Membrane Proteins.J Phys Chem B. 2017 Jun 1;121(21):5312-5321. doi: 10.1021/acs.jpcb.7b01705. Epub 2017 May 23. J Phys Chem B. 2017. PMID: 28485937
-
Characterization of Bifunctional Spin Labels for Investigating the Structural and Dynamic Properties of Membrane Proteins Using EPR Spectroscopy.J Phys Chem B. 2017 Oct 5;121(39):9185-9195. doi: 10.1021/acs.jpcb.7b07631. Epub 2017 Sep 26. J Phys Chem B. 2017. PMID: 28877443
-
Role of membrane mimetics on biophysical EPR studies of membrane proteins.Biochim Biophys Acta Biomembr. 2023 Apr;1865(4):184138. doi: 10.1016/j.bbamem.2023.184138. Epub 2023 Feb 9. Biochim Biophys Acta Biomembr. 2023. PMID: 36764474 Review.
-
Electron Paramagnetic Resonance as a Tool for Studying Membrane Proteins.Biomolecules. 2020 May 13;10(5):763. doi: 10.3390/biom10050763. Biomolecules. 2020. PMID: 32414134 Free PMC article. Review.
Cited by
-
Comparing the structural dynamics of the human KCNE3 in reconstituted micelle and lipid bilayered vesicle environments.Biochim Biophys Acta Biomembr. 2022 Oct 1;1864(10):183974. doi: 10.1016/j.bbamem.2022.183974. Epub 2022 Jun 15. Biochim Biophys Acta Biomembr. 2022. PMID: 35716725 Free PMC article.
-
Perspective on the Effect of Membrane Mimetics on Dynamic Properties of Integral Membrane Proteins.J Phys Chem B. 2023 May 4;127(17):3757-3765. doi: 10.1021/acs.jpcb.2c07324. Epub 2023 Apr 20. J Phys Chem B. 2023. PMID: 37078594 Free PMC article. Review.
-
Electron paramagnetic resonance spectroscopic characterization of the human KCNE3 protein in lipodisq nanoparticles for structural dynamics of membrane proteins.Biophys Chem. 2023 Oct;301:107080. doi: 10.1016/j.bpc.2023.107080. Epub 2023 Jul 26. Biophys Chem. 2023. PMID: 37531799 Free PMC article.
-
Synthesis, Characterization, and Nanodisc Formation of Non-ionic Polymers*.Angew Chem Int Ed Engl. 2021 Jul 26;60(31):16885-16888. doi: 10.1002/anie.202101950. Epub 2021 Jun 28. Angew Chem Int Ed Engl. 2021. PMID: 33998111 Free PMC article.
-
Lipid Membrane Mimetics in Functional and Structural Studies of Integral Membrane Proteins.Membranes (Basel). 2021 Sep 3;11(9):685. doi: 10.3390/membranes11090685. Membranes (Basel). 2021. PMID: 34564502 Free PMC article. Review.
References
-
- Overington JP; Al-Lazikani B; Hopkins AL How many drug targets are there? Nat. Rev. Drug Discovery 2006, 5, 993–996. - PubMed
-
- Sanders CR; Myers JK Disease-related misassembly of membrane proteins. Annu. Rev. Biophys. Biomol. Struct 2004, 33, 25–51. - PubMed
-
- Hubbell WL; McHaourab HS; Altenbach C; Lietzow MA Watching proteins move using site-directed spin labeling. Structure 1996, 4, 779–783. - PubMed
-
- Klug CS; Feix JB Methods and applications of site-directed spin labeling EPR spectroscopy. Methods Cell Biol 2008, 84, 617–658. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

