Ab initio folding of a trefoil-fold motif reveals structural similarity with a β-propeller blade motif
- PMID: 32142181
- PMCID: PMC7184783
- DOI: 10.1002/pro.3850
Ab initio folding of a trefoil-fold motif reveals structural similarity with a β-propeller blade motif
Abstract
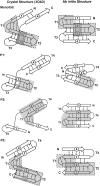
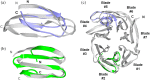
Many protein architectures exhibit evidence of internal rotational symmetry postulated to be the result of gene duplication/fusion events involving a primordial polypeptide motif. A common feature of such structures is a domain-swapped arrangement at the interface of the N- and C-termini motifs and postulated to provide cooperative interactions that promote folding and stability. De novo designed symmetric protein architectures have demonstrated an ability to accommodate circular permutation of the N- and C-termini in the overall architecture; however, the folding requirement of the primordial motif is poorly understood, and tolerance to circular permutation is essentially unknown. The β-trefoil protein fold is a threefold-symmetric architecture where the repeating ~42-mer "trefoil-fold" motif assembles via a domain-swapped arrangement. The trefoil-fold structure in isolation exposes considerable hydrophobic area that is otherwise buried in the intact β-trefoil trimeric assembly. The trefoil-fold sequence is not predicted to adopt the trefoil-fold architecture in ab initio folding studies; rather, the predicted fold is closely related to a compact "blade" motif from the β-propeller architecture. Expression of a trefoil-fold sequence and circular permutants shows that only the wild-type N-terminal motif definition yields an intact β-trefoil trimeric assembly, while permutants yield monomers. The results elucidate the folding requirements of the primordial trefoil-fold motif, and also suggest that this motif may sample a compact conformation that limits hydrophobic residue exposure, contains key trefoil-fold structural features, but is more structurally homologous to a β-propeller blade motif.
Keywords: domain swapping; folding pathway; protein evolution; protein symmetry.
© 2020 The Protein Society.
Conflict of interest statement
M.B. is a cofounder and has equity ownership in Trefoil Therapeutics, Inc.
Figures



References
-
- Lang D, Thoma R, Henn‐Sax M, Sterner R, Wilmanns M. Structural evidence for evolution of the beta/alpha barrel scaffold by gene duplication and fusion. Science. 2000;289:1546–1550. - PubMed
-
- Richter M, Bosnali M, Carstensen L, et al. Computational and experimental evidence for the evolution of a (βα)8‐barrel protein from an ancestral quarter‐barrel stabilized by disulfide bonds. J Mol Biol. 2010;398:763–773. - PubMed
-
- Fülöp V, Jones DT. β propellers: Structural rigidity and functional diversity. Curr Opin Struct Biol. 1999;9:715–721. - PubMed
-
- Chaudhuri I, Soding J, Lupas AN. Evolution of the β‐propeller fold. Proteins. 2008;71:795–803. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

