Crossover Position Drives Chromosome Remodeling for Accurate Meiotic Chromosome Segregation
- PMID: 32142707
- PMCID: PMC7162695
- DOI: 10.1016/j.cub.2020.01.079
Crossover Position Drives Chromosome Remodeling for Accurate Meiotic Chromosome Segregation
Abstract
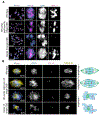
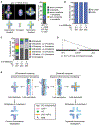
Interhomolog crossovers (COs) are a prerequisite for achieving accurate chromosome segregation during meiosis [1, 2]. COs are not randomly positioned, occurring at distinct genomic intervals during meiosis in all species examined [3-10]. The role of CO position as a major determinant of accurate chromosome segregation has not been previously directly analyzed in a metazoan. Here, we use spo-11 mutants, which lack endogenous DNA double-strand breaks (DSBs), to induce a single DSB by Mos1 transposon excision at defined chromosomal locations in the C. elegans germline and show that the position of the resulting CO directly affects the formation of distinct chromosome subdomains during meiotic chromosome remodeling. CO formation in the typically CO-deprived center region of autosomes leads to premature loss of sister chromatid cohesion and chromosome missegregation, whereas COs at an off-centered position, as in wild type, can result in normal remodeling and accurate segregation. Ionizing radiation (IR)-induced DSBs lead to the same outcomes, and modeling of IR dose-response reveals that the CO-unfavorable center region encompasses up to 6% of the total chromosome length. DSBs proximal to telomeres rarely form COs, likely because of formation of unstable recombination intermediates that cannot be sustained as chiasmata until late prophase. Our work supports a model in which regulation of CO position early in meiotic prophase is required for proper designation of chromosome subdomains and normal chromosome remodeling in late meiotic prophase I, resulting in accurate chromosome segregation and providing a mechanism to prevent aneuploid gamete formation.
Keywords: C. elegans; DNA double-strand break; DNA repair; Mos1; chromosome remodeling; chromosome segregation; crossover; germline; meiosis.
Copyright © 2020 Elsevier Ltd. All rights reserved.
Conflict of interest statement
Declaration of Interests The authors declare no competing interests.
Figures


Comment in
-
Meiosis: Location, Location, Location, How Crossovers Ensure Segregation.Curr Biol. 2020 Apr 6;30(7):R311-R313. doi: 10.1016/j.cub.2020.02.020. Curr Biol. 2020. PMID: 32259504
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

