Comparative differential cuproproteomes of Rhodobacter capsulatus reveal novel copper homeostasis related proteins
- PMID: 32149296
- PMCID: PMC7192791
- DOI: 10.1039/c9mt00314b
Comparative differential cuproproteomes of Rhodobacter capsulatus reveal novel copper homeostasis related proteins
Abstract
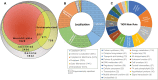
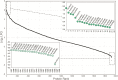
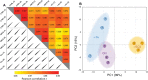
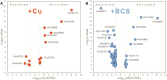
Copper (Cu) is an essential, but toxic, micronutrient for living organisms and cells have developed sophisticated response mechanisms towards both the lack and the excess of Cu in their environments. In this study, we achieved a global view of Cu-responsive changes in the prokaryotic model organism Rhodobacter capsulatus using label-free quantitative differential proteomics. Semi-aerobically grown cells under heterotrophic conditions in minimal medium (∼0.3 μM Cu) were compared with cells supplemented with either 5 μM Cu or with 5 mM of the Cu-chelator bathocuproine sulfonate. Mass spectrometry based bottom-up proteomics of unfractionated cell lysates identified 2430 of the 3632 putative proteins encoded by the genome, producing a robust proteome dataset for R. capsulatus. Use of biological and technical replicates for each growth condition yielded high reproducibility and reliable quantification for 1926 of the identified proteins. Comparison of cells grown under Cu-excess or Cu-depleted conditions to those grown under minimal Cu-sufficient conditions revealed that 75 proteins exhibited statistically significant (p < 0.05) abundance changes, ranging from 2- to 300-fold. A subset of the highly Cu-responsive proteins was orthogonally probed using molecular genetics, validating that several of them were indeed involved in cellular Cu homeostasis.
Figures




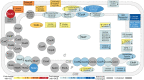
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

