Long transposon-rich centromeres in an oomycete reveal divergence of centromere features in Stramenopila-Alveolata-Rhizaria lineages
- PMID: 32150559
- PMCID: PMC7082073
- DOI: 10.1371/journal.pgen.1008646
Long transposon-rich centromeres in an oomycete reveal divergence of centromere features in Stramenopila-Alveolata-Rhizaria lineages
Abstract
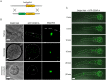
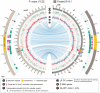
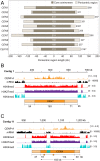
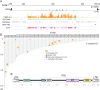
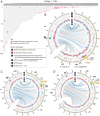
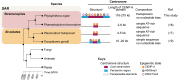
Centromeres are chromosomal regions that serve as platforms for kinetochore assembly and spindle attachments, ensuring accurate chromosome segregation during cell division. Despite functional conservation, centromere DNA sequences are diverse and often repetitive, making them challenging to assemble and identify. Here, we describe centromeres in an oomycete Phytophthora sojae by combining long-read sequencing-based genome assembly and chromatin immunoprecipitation for the centromeric histone CENP-A followed by high-throughput sequencing (ChIP-seq). P. sojae centromeres cluster at a single focus at different life stages and during nuclear division. We report an improved genome assembly of the P. sojae reference strain, which enabled identification of 15 enriched CENP-A binding regions as putative centromeres. By focusing on a subset of these regions, we demonstrate that centromeres in P. sojae are regional, spanning 211 to 356 kb. Most of these regions are transposon-rich, poorly transcribed, and lack the histone modification H3K4me2 but are embedded within regions with the heterochromatin marks H3K9me3 and H3K27me3. Strikingly, we discovered a Copia-like transposon (CoLT) that is highly enriched in the CENP-A chromatin. Similar clustered elements are also found in oomycete relatives of P. sojae, and may be applied as a criterion for prediction of oomycete centromeres. This work reveals a divergence of centromere features in oomycetes as compared to other organisms in the Stramenopila-Alveolata-Rhizaria (SAR) supergroup including diatoms and Plasmodium falciparum that have relatively short and simple regional centromeres. Identification of P. sojae centromeres in turn also advances the genome assembly.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

