New Insights into Modelling Bacterial Growth with Reference to the Fish Pathogen Flavobacterium psychrophilum
- PMID: 32150847
- PMCID: PMC7143051
- DOI: 10.3390/ani10030435
New Insights into Modelling Bacterial Growth with Reference to the Fish Pathogen Flavobacterium psychrophilum
Abstract
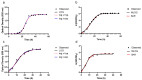
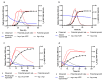
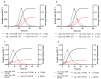
Two new models, based upon the principles promulgated by Baranyi and co-workers are presented and resulting growth functions evaluated based upon their ability to mimic bacterial growth of the fish pathogen Flavobacterium psychrophilum. These growth functions make use of a dampening function to suppress potential growth, represented by a logistic, and are derived from rate:state differential equations. Dampening effects are represented by a rectangular hyperbola or a simple exponential, incorporated into a logistic differential equation and solved analytically resulting in two newly derived growth equations, viz. logistic × hyperbola (log × hyp) and logistic × exponential (log × exp). These characteristics result in flexible and robust growth functions that can be expressed as equations with biologically meaningful parameters. The newly derived functions (log × hyp and log × exp), along with the Baranyi (BAR), simple logistic (LOG) and its modified form (MLOG) were evaluated based upon examination of residuals and measures of goodness-of-fit and cross-validation. Using these criteria, log × hyp, log × exp and BAR performed better than, or at least equally well as, LOG and MLOG. In contrast with log × exp and BAR, log × hyp can be easily manipulated mathematically allowing for simple algebraic expressions for time and microbial biomass at inflexion point, in addition to maximum and scaled maximum growth rates.
Keywords: Flavobacterium psychrophilum; bacterial diseases; farmed fish; modelling.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Figures
References
-
- Whiting R.C. Notes on reparameterization of bacterial growth curves—A reply to J. Baranyi and W. E. Garthright. Food Microbiol. 1992;9:173–174. doi: 10.1016/0740-0020(92)80026-Z. - DOI
Grants and funding
LinkOut - more resources
Full Text Sources

