New Perspectives Related to the Bioluminescent System in Dinoflagellates: Pyrocystis lunula, a Case Study
- PMID: 32150894
- PMCID: PMC7084563
- DOI: 10.3390/ijms21051784
New Perspectives Related to the Bioluminescent System in Dinoflagellates: Pyrocystis lunula, a Case Study
Abstract
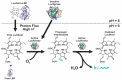
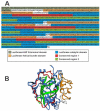
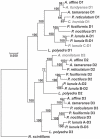
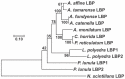
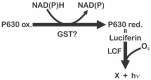
Pyrocystis lunula is considered a model organism due to its bioluminescence capacity linked to circadian rhythms. The mechanisms underlying the bioluminescent phenomenon have been well characterized in dinoflagellates; however, there are still some aspects that remain an enigma. Such is the case of the presence and diversity of the luciferin-binding protein (LBP), as well as the synthesis process of luciferin. Here we carry out a review of the literature in relation to the molecular players responsible for bioluminescence in dinoflagellates, with particular interest in P. lunula. We also carried out a phylogenetic analysis of the conservation of protein sequence, structure and evolutionary pattern of these key players. The basic structure of the luciferase (LCF) is quite conserved among the sequences reported to date for dinoflagellate species, but not in the case of the LBP, which has proven to be more variable in terms of sequence and structure. In the case of luciferin, its synthesis has been shown to be complex process with more than one metabolic pathway involved. The glutathione S-transferase (GST) and the P630 or blue compound, seem to be involved in this process. In the same way, various hypotheses regarding the role of bioluminescence in dinoflagellates are exposed.
Keywords: P630; blue compound; glutathione S-transferase; luciferase; luciferin; luciferin-binding protein.
Conflict of interest statement
The authors declare no conflict of interest.
Figures






References
-
- Widder E. Marine bioluminescence. Biosci. Explain. 2001;1:1–9.
-
- Shimomura O. Bioluminescence: Chemical Principles and Methods. World Scientific Publishing Co. Pte. Ltd.; Singapore: 2012. Revised Edition.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

