Using Genetic Distance from Archived Samples for the Prediction of Antibiotic Resistance in Escherichia coli
- PMID: 32152083
- PMCID: PMC7179619
- DOI: 10.1128/AAC.02417-19
Using Genetic Distance from Archived Samples for the Prediction of Antibiotic Resistance in Escherichia coli
Abstract
The rising rates of antibiotic resistance increasingly compromise empirical treatment. Knowing the antibiotic susceptibility of a pathogen's close genetic relative(s) may improve empirical antibiotic selection. Using genomic and phenotypic data for Escherichia coli isolates from three separate clinically derived databases, we evaluated multiple genomic methods and statistical models for predicting antibiotic susceptibility, focusing on potentially rapidly available information, such as lineage or genetic distance from archived isolates. We applied these methods to derive and validate the prediction of antibiotic susceptibility to common antibiotics. We evaluated 968 separate episodes of suspected and confirmed infection with Escherichia coli from three geographically and temporally separated databases in Ontario, Canada, from 2010 to 2018. Across all approaches, model performance (area under the curve [AUC]) ranges for predicting antibiotic susceptibility were the greatest for ciprofloxacin (AUC, 0.76 to 0.97) and the lowest for trimethoprim-sulfamethoxazole (AUC, 0.51 to 0.80). When a model predicted that an isolate was susceptible, the resulting (posttest) probabilities of susceptibility were sufficient to warrant empirical therapy for most antibiotics (mean, 92%). An approach combining multiple models could permit the use of narrower-spectrum oral agents in 2 out of every 3 patients while maintaining high treatment adequacy (∼90%). Methods based on genetic relatedness to archived samples of E. coli could be used to predict antibiotic resistance and improve antibiotic selection.
Keywords: Gram-negative bacteria; antibiotic-resistant organisms; antibiotics; empirical antibiotics; genomics; rapid diagnostics.
Copyright © 2020 MacFadden et al.
Figures


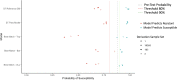

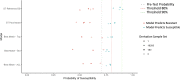
References
-
- Review on Antimicrobial Resistance. 2016. Tackling drug-resistant infections globally: final report and recommendations. Her Majesty’s Government, London, United Kingdom.
-
- Frieden T. 2013. Antibiotic resistance threats in the United States 2013. Centers for Disease Control and Prevention, Atlanta, GA.
-
- Kumar A, Roberts D, Wood KE, Light B, Parrillo JE, Sharma S, Suppes R, Feinstein D, Zanotti S, Taiberg L, Gurka D, Kumar A, Cheang M. 2006. Duration of hypotension before initiation of effective antimicrobial therapy is the critical determinant of survival in human septic shock. Crit Care Med 34:1589–1596. doi: 10.1097/01.CCM.0000217961.75225.E9. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

