Fluorescence-Reported Allelic Exchange Mutagenesis-Mediated Gene Deletion Indicates a Requirement for Chlamydia trachomatis Tarp during In Vivo Infectivity and Reveals a Specific Role for the C Terminus during Cellular Invasion
- PMID: 32152196
- PMCID: PMC7171248
- DOI: 10.1128/IAI.00841-19
Fluorescence-Reported Allelic Exchange Mutagenesis-Mediated Gene Deletion Indicates a Requirement for Chlamydia trachomatis Tarp during In Vivo Infectivity and Reveals a Specific Role for the C Terminus during Cellular Invasion
Abstract
The translocated actin recruiting phosphoprotein (Tarp) is a multidomain type III secreted effector used by Chlamydia trachomatis In aggregate, existing data suggest a role of this effector in initiating new infections. As new genetic tools began to emerge to study chlamydial genes in vivo, we speculated as to what degree Tarp function contributes to Chlamydia's ability to parasitize mammalian host cells. To address this question, we generated a complete tarP deletion mutant using the fluorescence-reported allelic exchange mutagenesis (FRAEM) technique and complemented the mutant in trans with wild-type tarP or mutant tarP alleles engineered to harbor in-frame domain deletions. We provide evidence for the significant role of Tarp in C. trachomatis invasion of host cells. Complementation studies indicate that the C-terminal filamentous actin (F-actin)-binding domains are responsible for Tarp-mediated invasion efficiency. Wild-type C. trachomatis entry into HeLa cells resulted in host cell shape changes, whereas the tarP mutant did not. Finally, using a novel cis complementation approach, C. trachomatis lacking tarP demonstrated significant attenuation in a murine genital tract infection model. Together, these data provide definitive genetic evidence for the critical role of the Tarp F-actin-binding domains in host cell invasion and for the Tarp effector as a bona fide C. trachomatis virulence factor.
Keywords: Chlamydia trachomatis; Tarp; actin; actin cytoskeleton; cytoskeleton; effector; effector functions.
Copyright © 2020 American Society for Microbiology.
Figures



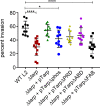


References
-
- Moulder JW, Hatch TP, Kuo CC, Schachter J, Storz J. 1984. Chlamydia, p 729–739. In Krieg NR. (ed), Bergey’s manual of systematic bacteriology. Williams & Wilkins, Baltimore, MD.
-
- Clifton DR, Fields KA, Grieshaber SS, Dooley CA, Fischer ER, Mead DJ, Carabeo RA, Hackstadt T. 2004. A chlamydial type III translocated protein is tyrosine-phosphorylated at the site of entry and associated with recruitment of actin. Proc Natl Acad Sci U S A 101:10166–10171. doi:10.1073/pnas.0402829101. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

