A widespread role for SLC transmembrane transporters in resistance to cytotoxic drugs
- PMID: 32152546
- PMCID: PMC7610918
- DOI: 10.1038/s41589-020-0483-3
A widespread role for SLC transmembrane transporters in resistance to cytotoxic drugs
Abstract
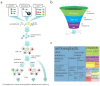
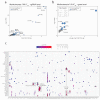
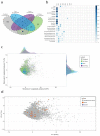
Solute carriers (SLCs) are the largest family of transmembrane transporters in humans and are major determinants of cellular metabolism. Several SLCs have been shown to be required for the uptake of chemical compounds into cellular systems, but systematic surveys of transporter-drug relationships in human cells are currently lacking. We performed a series of genetic screens in a haploid human cell line against 60 cytotoxic compounds representative of the chemical space populated by approved drugs. By using an SLC-focused CRISPR-Cas9 library, we identified transporters whose absence induced resistance to the drugs tested. This included dependencies involving the transporters SLC11A2/SLC16A1 for artemisinin derivatives and SLC35A2/SLC38A5 for cisplatin. The functional dependence on SLCs observed for a significant proportion of the screened compounds suggests a widespread role for SLCs in the uptake and cellular activity of cytotoxic drugs and provides an experimentally validated set of SLC-drug associations for a number of clinically relevant compounds.
Conflict of interest statement
The authors declare no competing financial interests. C-H. L. is an employee of AstraZeneca Limited (UK).
Figures


Comment in
-
Hitchhiking into the cell.Nat Chem Biol. 2020 Apr;16(4):367-368. doi: 10.1038/s41589-020-0489-x. Nat Chem Biol. 2020. PMID: 32152545 No abstract available.
References
-
- Gonzalez FJ, Coughtrie M, Tukey RH. Drug Metabolism. In: Brunton LL, Chabner BA, Knollmann BC, editors. Goodman & Gilman’s: The Pharmacological Basis of Therapeutics. McGraw-Hill Education; 2015.
-
- César-Razquin A, et al. A Call for Systematic Research on Solute Carriers. Cell. 2015;162:478–487. - PubMed
-
- Fletcher JI, Haber M, Henderson MJ, Norris MD. ABC transporters in cancer: more than just drug efflux pumps. Nat Rev Cancer. 2010;10:147–156. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

