Genetic diversity and evolutionary analysis of human respirovirus type 3 strains isolated in Kenya using complete hemagglutinin-neuraminidase (HN) gene
- PMID: 32155160
- PMCID: PMC7064169
- DOI: 10.1371/journal.pone.0229355
Genetic diversity and evolutionary analysis of human respirovirus type 3 strains isolated in Kenya using complete hemagglutinin-neuraminidase (HN) gene
Abstract
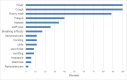
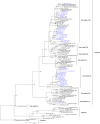
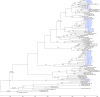
Human respirovirus type 3 (HRV3) is a leading etiology of lower respiratory tract infections in young children and ranks only second to the human respiratory syncytial virus (HRSV). Despite the public health importance of HRV3, there is limited information about the genetic characteristics and diversity of these viruses in Kenya. To begin to address this gap, we analyzed 35 complete hemagglutinin-neuraminidase (HN) sequences of HRV3 strains isolated in Kenya between 2010 and 2013. Viral RNA was extracted from the isolates, and the entire HN gene amplified by RT-PCR followed by nucleotide sequencing. Phylogenetic analyses of the sequences revealed that all the Kenyan isolates grouped into genetic Cluster C; sub-clusters C1a, C2, and C3a. The majority (54%) of isolates belonged to sub-cluster C3a, followed by C2 (43%) and C1a (2.9%). Sequence analysis revealed high identities between the Kenyan isolates and the HRV3 prototype strain both at the amino acid (96.5-97.9%) and nucleotide (94.3-95.6%) levels. No amino acid variations affecting the catalytic/active sites of the HN glycoprotein were observed among the Kenyan isolates. Selection pressure analyses showed that the HN glycoprotein was evolving under positive selection. Evolutionary analyses revealed that the mean TMRCA for the HN sequence dataset was 1942 (95% HPD: 1928-1957), while the mean evolutionary rate was 4.65x10-4 nucleotide substitutions/site/year (95% HPD: 2.99x10-4 to 6.35x10-4). Overall, our results demonstrate the co-circulation of strains of cluster C HRV3 variants in Kenya during the study period. This is the first study to describe the genetic and molecular evolutionary aspects of HRV3 in Kenya using the complete HN gene.
Conflict of interest statement
The authors have declared that no competing interests exist
Figures
References
-
- Smielewska A, Emmott E, Ranellou K, Popay A, Goodfellow I, Jalal H. UK circulating strains of human parainfluenza 3: An amplicon based next generation sequencing method and phylogenetic analysis [version 2; referees: 2 approved]. Wellcome Open Research. 2018. 10.12688/wellcomeopenres.14730.2 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

