Synthetic Lethality Screening Identifies FDA-Approved Drugs that Overcome ATP7B-Mediated Tolerance of Tumor Cells to Cisplatin
- PMID: 32155756
- PMCID: PMC7139527
- DOI: 10.3390/cancers12030608
Synthetic Lethality Screening Identifies FDA-Approved Drugs that Overcome ATP7B-Mediated Tolerance of Tumor Cells to Cisplatin
Abstract
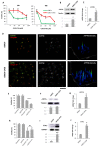
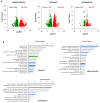
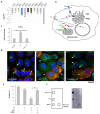
Tumor resistance to chemotherapy represents an important challenge in modern oncology. Although platinum (Pt)-based drugs have demonstrated excellent therapeutic potential, their effectiveness in a wide range of tumors is limited by the development of resistance mechanisms. One of these mechanisms includes increased cisplatin sequestration/efflux by the copper-transporting ATPase, ATP7B. However, targeting ATP7B to reduce Pt tolerance in tumors could represent a serious risk because suppression of ATP7B might compromise copper homeostasis, as happens in Wilson disease. To circumvent ATP7B-mediated Pt tolerance we employed a high-throughput screen (HTS) of an FDA/EMA-approved drug library to detect safe therapeutic molecules that promote cisplatin toxicity in the IGROV-CP20 ovarian carcinoma cells, whose resistance significantly relies on ATP7B. Using a synthetic lethality approach, we identified and validated three hits (Tranilast, Telmisartan, and Amphotericin B) that reduced cisplatin resistance. All three drugs induced Pt-mediated DNA damage and inhibited either expression or trafficking of ATP7B in a tumor-specific manner. Global transcriptome analyses showed that Tranilast and Amphotericin B affect expression of genes operating in several pathways that confer tolerance to cisplatin. In the case of Tranilast, these comprised key Pt-transporting proteins, including ATOX1, whose suppression affected ability of ATP7B to traffic in response to cisplatin. In summary, our findings reveal Tranilast, Telmisartan, and Amphotericin B as effective drugs that selectively promote cisplatin toxicity in Pt-resistant ovarian cancer cells and underscore the efficiency of HTS strategy for identification of biosafe compounds, which might be rapidly repurposed to overcome resistance of tumors to Pt-based chemotherapy.
Keywords: ATP7B; FDA-approved drugs; cancer; cisplatin resistance; copper transporters; synthetic lethality screening.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



Similar articles
-
TFEB Regulates ATP7B Expression to Promote Platinum Chemoresistance in Human Ovarian Cancer Cells.Cells. 2022 Jan 10;11(2):219. doi: 10.3390/cells11020219. Cells. 2022. PMID: 35053335 Free PMC article.
-
Modulation of the cellular pharmacology of JM118, the major metabolite of satraplatin, by copper influx and efflux transporters.Cancer Chemother Pharmacol. 2006 Jun;57(6):781-8. doi: 10.1007/s00280-005-0121-5. Epub 2005 Sep 17. Cancer Chemother Pharmacol. 2006. PMID: 16170571
-
Mechanism of tumor resistance to cisplatin mediated by the copper transporter ATP7B.Biochem Cell Biol. 2011 Apr;89(2):138-47. doi: 10.1139/o10-150. Biochem Cell Biol. 2011. PMID: 21455266 Review.
-
Prognostic value of the Cu-transporting ATPase in ovarian carcinoma patients receiving cisplatin-based chemotherapy.Clin Cancer Res. 2004 Apr 15;10(8):2804-11. doi: 10.1158/1078-0432.ccr-03-0454. Clin Cancer Res. 2004. PMID: 15102688
-
Role of copper transporters in platinum resistance.World J Clin Oncol. 2016 Feb 10;7(1):106-13. doi: 10.5306/wjco.v7.i1.106. World J Clin Oncol. 2016. PMID: 26862494 Free PMC article. Review.
Cited by
-
The 'speck'-tacular oversight of the NLRP3-pyroptosis pathway on gastrointestinal inflammatory diseases and tumorigenesis.J Biomed Sci. 2023 Oct 27;30(1):90. doi: 10.1186/s12929-023-00983-7. J Biomed Sci. 2023. PMID: 37891577 Free PMC article. Review.
-
Unveiling the genomic landscape of possible metastatic malignant transformation of teratoma secondary to cisplatin-chemotherapy: a Tempus gene analysis-based case report literature review.Front Oncol. 2023 Jun 22;13:1192843. doi: 10.3389/fonc.2023.1192843. eCollection 2023. Front Oncol. 2023. PMID: 37427132 Free PMC article.
-
Activation of Adrenoceptor Alpha-2 (ADRA2A) Promotes Chemosensitization to Carboplatin in Ovarian Cancer Cell Lines.Curr Issues Mol Biol. 2023 Nov 28;45(12):9566-9578. doi: 10.3390/cimb45120598. Curr Issues Mol Biol. 2023. PMID: 38132444 Free PMC article.
-
Molecular mechanisms of cisplatin resistance in ovarian cancer.Genes Dis. 2023 Aug 2;11(6):101063. doi: 10.1016/j.gendis.2023.06.032. eCollection 2024 Nov. Genes Dis. 2023. PMID: 39224110 Free PMC article. Review.
-
Patient-derived tumor xenograft animal model of gastric cancer in precision chemotherapy.Naunyn Schmiedebergs Arch Pharmacol. 2025 Aug;398(8):10305-10316. doi: 10.1007/s00210-025-03903-8. Epub 2025 Feb 19. Naunyn Schmiedebergs Arch Pharmacol. 2025. PMID: 39969603
References
-
- Lee Y.Y., Choi C.H., Do I.G., Song S.Y., Lee W., Park H.S., Song T.J., Kim M.K., Kim T.J., Lee J.W., et al. Prognostic value of the copper transporters, CTR1 and CTR2, in patients with ovarian carcinoma receiving platinum-based chemotherapy. Gynecol. Oncol. 2011;122:361–365. doi: 10.1016/j.ygyno.2011.04.025. - DOI - PubMed
-
- Komatsu M., Sumizawa T., Mutoh M., Chen Z.S., Terada K., Furukawa T., Yang X.L., Gao H., Miura N., Sugiyama T., et al. Copper-transporting P-type adenosine triphosphatase (ATP7B) is associated with cisplatin resistance. Cancer Res. 2000;60:1312–1316. - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

