Probing the Transition State in Enzyme Catalysis by High-Pressure NMR Dynamics
- PMID: 32159076
- PMCID: PMC7063682
- DOI: 10.1038/s41929-019-0307-6
Probing the Transition State in Enzyme Catalysis by High-Pressure NMR Dynamics
Abstract
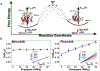
Protein conformational changes are frequently essential for enzyme catalysis, and in several cases, shown to be the limiting factor for overall catalytic speed. However, a structural understanding of corresponding transition states, needed to rationalize the kinetics, remains obscure due to their fleeting nature. Here, we determine the transition-state ensemble of the rate-limiting conformational transition in the enzyme adenylate kinase, by a synergistic approach between experimental high-pressure NMR relaxation during catalysis and molecular dynamics simulations. By comparing homologous kinases evolved under ambient or high pressure in the deep-sea, we detail transition state ensembles that differ in solvation as directly measured by the pressure dependence of catalysis. Capturing transition-state ensembles begins to complete the catalytic energy landscape that is generally characterized by structures of all intermediates and frequencies of transitions among them.
Conflict of interest statement
Competing Interests The authors declare no competing interests.
Figures
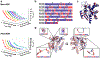


Similar articles
-
Structural basis for ligand binding to an enzyme by a conformational selection pathway.Proc Natl Acad Sci U S A. 2017 Jun 13;114(24):6298-6303. doi: 10.1073/pnas.1700919114. Epub 2017 May 30. Proc Natl Acad Sci U S A. 2017. PMID: 28559350 Free PMC article.
-
Wide transition-state ensemble as key component for enzyme catalysis.Elife. 2025 Feb 18;12:RP93099. doi: 10.7554/eLife.93099. Elife. 2025. PMID: 39963964 Free PMC article.
-
Transition-state ensemble in enzyme catalysis: possibility, reality, or necessity?J Theor Biol. 2000 Apr 21;203(4):383-97. doi: 10.1006/jtbi.2000.1097. J Theor Biol. 2000. PMID: 10736215
-
Probing invisible, low-populated States of protein molecules by relaxation dispersion NMR spectroscopy: an application to protein folding.Acc Chem Res. 2008 Mar;41(3):442-51. doi: 10.1021/ar700189y. Epub 2008 Feb 15. Acc Chem Res. 2008. PMID: 18275162 Review.
-
Folding funnels and conformational transitions via hinge-bending motions.Cell Biochem Biophys. 1999;31(2):141-64. doi: 10.1007/BF02738169. Cell Biochem Biophys. 1999. PMID: 10593256 Review.
Cited by
-
Adaptations for Pressure and Temperature in Dihydrofolate Reductases.Microorganisms. 2021 Aug 11;9(8):1706. doi: 10.3390/microorganisms9081706. Microorganisms. 2021. PMID: 34442785 Free PMC article.
-
Evolutionary-Scale Enzymology Enables Biochemical Constant Prediction Across a Multi-Peaked Catalytic Landscape.bioRxiv [Preprint]. 2024 Oct 25:2024.10.23.619915. doi: 10.1101/2024.10.23.619915. bioRxiv. 2024. PMID: 39484523 Free PMC article. Preprint.
-
High Pressure CPMG and CEST Reveal That Cavity Position Dictates Distinct Dynamic Disorder in the PP32 Repeat Protein.J Phys Chem B. 2022 Dec 22;126(50):10597-10607. doi: 10.1021/acs.jpcb.2c05498. Epub 2022 Dec 1. J Phys Chem B. 2022. PMID: 36455152 Free PMC article.
-
NMR in the Age of Modern Biomedical Research and Drug Discovery.J Mol Biol. 2025 Jun 23:169302. doi: 10.1016/j.jmb.2025.169302. Online ahead of print. J Mol Biol. 2025. PMID: 40562354 Review.
-
Switching Go̅-Martini for Investigating Protein Conformational Transitions and Associated Protein-Lipid Interactions.J Chem Theory Comput. 2024 Mar 26;20(6):2618-2629. doi: 10.1021/acs.jctc.3c01222. Epub 2024 Mar 6. J Chem Theory Comput. 2024. PMID: 38447049 Free PMC article.
References
Grants and funding
LinkOut - more resources
Full Text Sources
