MKS-NPHP module proteins control ciliary shedding at the transition zone
- PMID: 32163404
- PMCID: PMC7093003
- DOI: 10.1371/journal.pbio.3000640
MKS-NPHP module proteins control ciliary shedding at the transition zone
Abstract
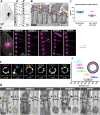
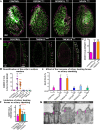
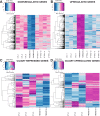
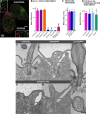
Ciliary shedding occurs from unicellular organisms to metazoans. Although required during the cell cycle and during neurogenesis, the process remains poorly understood. In all cellular models, this phenomenon occurs distal to the transition zone (TZ), suggesting conserved molecular mechanisms. The TZ module proteins (Meckel Gruber syndrome [MKS]/Nephronophtysis [NPHP]/Centrosomal protein of 290 kDa [CEP290]/Retinitis pigmentosa GTPase regulator-Interacting Protein 1-Like Protein [RPGRIP1L]) are known to cooperate to establish TZ formation and function. To determine whether they control deciliation, we studied the function of 5 of them (Transmembrane protein 107 [TMEM107], Transmembrane protein 216 [TMEM216], CEP290, RPGRIP1L, and NPHP4) in Paramecium. All proteins are recruited to the TZ of growing cilia and localize with 9-fold symmetry at the level of the most distal part of the TZ. We demonstrate that depletion of the MKS2/TMEM216 and TMEM107 proteins induces constant deciliation of some cilia, while depletion of either NPHP4, CEP290, or RPGRIP1L prevents Ca2+/EtOH deciliation. Our results constitute the first evidence for a role of conserved TZ proteins in deciliation and open new directions for understanding motile cilia physiology.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
References
-
- Huang L, Szymanska K, Jensen VL, Janecke AR, Innes AM, Davis EE, et al. TMEM237 is mutated in individuals with a Joubert syndrome related disorder and expands the role of the TMEM family at the ciliary transition zone. Am J Hum Genet. 2011. December 9;89(6):713–30. 10.1016/j.ajhg.2011.11.005 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

