The establishment of reference sequence for SARS-CoV-2 and variation analysis
- PMID: 32167180
- PMCID: PMC7228400
- DOI: 10.1002/jmv.25762
The establishment of reference sequence for SARS-CoV-2 and variation analysis
Abstract
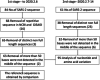
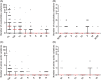
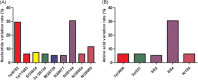
Starting around December 2019, an epidemic of pneumonia, which was named COVID-19 by the World Health Organization, broke out in Wuhan, China, and is spreading throughout the world. A new coronavirus, named severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) by the Coronavirus Study Group of the International Committee on Taxonomy of Viruses was soon found to be the cause. At present, the sensitivity of clinical nucleic acid detection is limited, and it is still unclear whether it is related to genetic variation. In this study, we retrieved 95 full-length genomic sequences of SARAS-CoV-2 strains from the National Center for Biotechnology Information and GISAID databases, established the reference sequence by conducting multiple sequence alignment and phylogenetic analyses, and analyzed sequence variations along the SARS-CoV-2 genome. The homology among all viral strains was generally high, among them, 99.99% (99.91%-100%) at the nucleotide level and 99.99% (99.79%-100%) at the amino acid level. Although overall variation in open-reading frame (ORF) regions is low, 13 variation sites in 1a, 1b, S, 3a, M, 8, and N regions were identified, among which positions nt28144 in ORF 8 and nt8782 in ORF 1a showed mutation rate of 30.53% (29/95) and 29.47% (28/95), respectively. These findings suggested that there may be selective mutations in SARS-COV-2, and it is necessary to avoid certain regions when designing primers and probes. Establishment of the reference sequence for SARS-CoV-2 could benefit not only biological study of this virus but also diagnosis, clinical monitoring and intervention of SARS-CoV-2 infection in the future.
Keywords: SARS-CoV-2; homology; nucleotide; reference sequence; variation.
© 2020 Wiley Periodicals, Inc.
Conflict of interest statement
The authors declare that there are no conflict of interests.
Figures
References
-
- National Health Commission of the People's Republic of China . http://www.nhc.gov.cn (Assessed on March 8th, 2020). - PMC - PubMed
-
- WHO main website . https://www.who.int (accessed March 8th, 2020).
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

