PNA-Based MicroRNA Detection Methodologies
- PMID: 32178411
- PMCID: PMC7144472
- DOI: 10.3390/molecules25061296
PNA-Based MicroRNA Detection Methodologies
Abstract
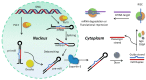
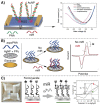
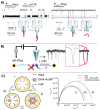
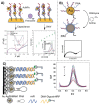
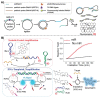
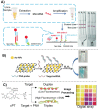
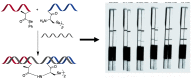
MicroRNAs (miRNAs or miRs) are small noncoding RNAs involved in the fine regulation of post-transcriptional processes in the cell. The physiological levels of these short (20-22-mer) oligonucleotides are important for the homeostasis of the organism, and therefore dysregulation can lead to the onset of cancer and other pathologies. Their importance as biomarkers is constantly growing and, in this context, detection methods based on the hybridization to peptide nucleic acids (PNAs) are gaining their place in the spotlight. After a brief overview of their biogenesis, this review will discuss the significance of targeting miR, providing a wide range of PNA-based approaches to detect them at biologically significant concentrations, based on electrochemical, fluorescence and colorimetric assays.
Keywords: colorimetric detection; electrochemical biosensors; fluorescence; light-triggered; microRNA; nanoparticles; peptide nucleic acid (PNA); templated reactions.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



Similar articles
-
Amplification-Free Detection of Circulating microRNA Biomarkers from Body Fluids Based on Fluorogenic Oligonucleotide-Templated Reaction between Engineered Peptide Nucleic Acid Probes: Application to Prostate Cancer Diagnosis.Anal Chem. 2016 Aug 16;88(16):8091-8. doi: 10.1021/acs.analchem.6b01594. Epub 2016 Aug 7. Anal Chem. 2016. PMID: 27498854
-
Chemical structure requirements and cellular targeting of microRNA-122 by peptide nucleic acids anti-miRs.Nucleic Acids Res. 2012 Mar;40(5):2152-67. doi: 10.1093/nar/gkr885. Epub 2011 Nov 8. Nucleic Acids Res. 2012. PMID: 22070883 Free PMC article.
-
miR-122 targeting with LNA/2'-O-methyl oligonucleotide mixmers, peptide nucleic acids (PNA), and PNA-peptide conjugates.RNA. 2008 Feb;14(2):336-46. doi: 10.1261/rna.844108. Epub 2007 Dec 11. RNA. 2008. PMID: 18073344 Free PMC article.
-
Peptide Nucleic Acid-Based Biosensors for Cancer Diagnosis.Molecules. 2017 Nov 11;22(11):1951. doi: 10.3390/molecules22111951. Molecules. 2017. PMID: 29137122 Free PMC article. Review.
-
Peptide nucleic acid (PNA) binding-mediated gene regulation.Cell Res. 2004 Apr;14(2):111-6. doi: 10.1038/sj.cr.7290209. Cell Res. 2004. PMID: 15115611 Review.
Cited by
-
Integrating high-performing electrochemical transducers in lateral flow assay.Anal Bioanal Chem. 2021 Sep;413(22):5535-5549. doi: 10.1007/s00216-021-03301-y. Epub 2021 Apr 28. Anal Bioanal Chem. 2021. PMID: 33913001 Free PMC article.
-
Investigating the Interactions of Peptide Nucleic Acids with Multicomponent Peptide Hydrogels for the Advancement of Healthcare Technologies.Gels. 2025 May 17;11(5):367. doi: 10.3390/gels11050367. Gels. 2025. PMID: 40422386 Free PMC article.
-
Biosensing circulating MicroRNAs in autoinflammatory skin diseases: Focus on Hidradenitis suppurativa.Front Genet. 2024 Apr 9;15:1383452. doi: 10.3389/fgene.2024.1383452. eCollection 2024. Front Genet. 2024. PMID: 38655054 Free PMC article.
-
Chemical approaches to discover the full potential of peptide nucleic acids in biomedical applications.Beilstein J Org Chem. 2021 Jul 19;17:1641-1688. doi: 10.3762/bjoc.17.116. eCollection 2021. Beilstein J Org Chem. 2021. PMID: 34367346 Free PMC article. Review.
-
Development of a His-Tag-mediated pull-down and quantification assay for G-quadruplex containing DNA sequences.RSC Chem Biol. 2024 Nov 28;6(1):56-64. doi: 10.1039/d4cb00185k. eCollection 2025 Jan 2. RSC Chem Biol. 2024. PMID: 39634055 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

