PETRA: Drug Engineering via Rigidity Analysis
- PMID: 32178472
- PMCID: PMC7144111
- DOI: 10.3390/molecules25061304
PETRA: Drug Engineering via Rigidity Analysis
Abstract
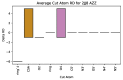
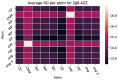
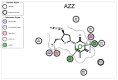
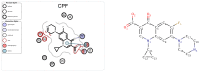
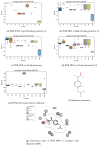
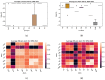
Rational drug design aims to develop pharmaceutical agents that impart maximal therapeutic benefits via their interaction with their intended biological targets. In the past several decades, advances in computational tools that inform wet-lab techniques have aided the development of a wide variety of new medicines with high efficacies. Nonetheless, drug development remains a time and cost intensive process. In this work, we have developed a computational pipeline for assessing how individual atoms contribute to a ligand's effect on the structural stability of a biological target. Our approach takes as input a protein-ligand resolved PDB structure file and systematically generates all possible ligand variants. We assess how the atomic-level edits to the ligand alter the drug's effect via a graph theoretic rigidity analysis approach. We demonstrate, via four case studies of common drugs, the utility of our pipeline and corroborate our analyses with known biophysical properties of the medicines, as reported in the literature.
Keywords: ligand engineering; rigidity analysis.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



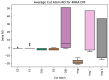
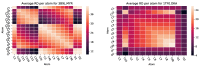
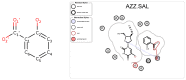

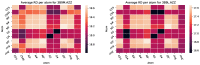
References
-
- Engh R.A., Brandstetter H., Sucher G., Eichinger A., Baumann U., Bode W., Huber R., Poll T., Rudolph R., von der Saal W. Enzyme flexibility, solvent and ‘weak’interactions characterize thrombin–ligand interactions: Implications for drug design. Structure. 1996;4:1353–1362. doi: 10.1016/S0969-2126(96)00142-6. - DOI - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

