Mitf Links Neuronal Activity and Long-Term Homeostatic Intrinsic Plasticity
- PMID: 32193365
- PMCID: PMC7174873
- DOI: 10.1523/ENEURO.0412-19.2020
Mitf Links Neuronal Activity and Long-Term Homeostatic Intrinsic Plasticity
Abstract
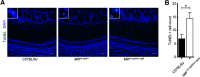
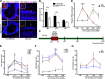
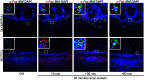
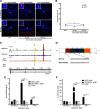
Neuroplasticity forms the basis for neuronal circuit complexity and differences between otherwise similar circuits. We show that the microphthalmia-associated transcription factor (Mitf) plays a central role in intrinsic plasticity of olfactory bulb (OB) projection neurons. Mitral and tufted (M/T) neurons from Mitf mutant mice are hyperexcitable, have a reduced A-type potassium current (IA) and exhibit reduced expression of Kcnd3, which encodes a potassium voltage-gated channel subunit (Kv4.3) important for generating the IA Furthermore, expression of the Mitf and Kcnd3 genes is activity dependent in OB projection neurons and the MITF protein activates expression from Kcnd3 regulatory elements. Moreover, Mitf mutant mice have changes in olfactory habituation and have increased habituation for an odorant following long-term exposure, indicating that regulation of Kcnd3 is pivotal for long-term olfactory adaptation. Our findings show that Mitf acts as a direct regulator of intrinsic homeostatic feedback and links neuronal activity, transcriptional changes and neuronal function.
Keywords: Kcnd3; genetics; hyperactivity; intrinsic plasticity; potassium channel; transcription.
Copyright © 2020 Atacho et al.
Figures






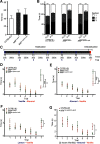

References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous
