A hybrid model of intercellular tension and cell-matrix mechanical interactions in a multicellular geometry
- PMID: 32193709
- PMCID: PMC7502553
- DOI: 10.1007/s10237-020-01321-8
A hybrid model of intercellular tension and cell-matrix mechanical interactions in a multicellular geometry
Abstract
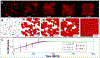
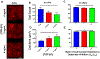
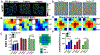
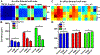
Epithelial cells form continuous sheets of cells that exist in tensional homeostasis. Homeostasis is maintained through cell-to-cell junctions that distribute tension and balance forces between cells and their underlying matrix. Disruption of tensional homeostasis can lead to epithelial-mesenchymal transition (EMT), a transdifferentiation process in which epithelial cells adopt a mesenchymal phenotype, losing cell-cell adhesion and enhancing cellular motility. This process is critical during embryogenesis and wound healing, but is also dysregulated in many disease states. To further understand the role of intercellular tension in spatial patterning of epithelial cell monolayers, we developed a multicellular computational model of cell-cell and cell-substrate forces. This work builds on a hybrid cellular Potts model (CPM)-finite element model to evaluate cell-matrix mechanical feedback of an adherent multicellular cluster. Cellular movement is governed by thermodynamic constraints from cell volume, cell-cell and cell-matrix contacts, and durotaxis, which arises from cell-generated traction forces on a finite element substrate. Junction forces at cell-cell contacts balance these traction forces, thereby producing a mechanically stable epithelial monolayer. Simulations were compared to in vitro experiments using fluorescence-based junction force sensors in clusters of cells undergoing EMT. Results indicate that the multicellular CPM model can reproduce many aspects of EMT, including epithelial monolayer formation dynamics, changes in cell geometry, and spatial patterning of cell-cell forces in an epithelial tissue.
Keywords: Cell mechanics; Cellular Potts model; Cell–cell junction forces; Epithelial–mesenchymal transition; Spatial patterning; Traction forces.
Figures



References
-
- Fristrom D, Tissue and Cell 20(5), 645 (1988). DOI 10.1016/0040-8166(88)90015-8. URL http://www.sciencedirect.com/science/article/pii/0040816688900158 - DOI - PubMed
-
- Ingber DE, Proceedings of the National Academy of Sciences 102(33), 11571 (2005). DOI 10.1073/pnas.0505939102. URL https://www.pnas.org/content/102/33/11571 - DOI - PMC - PubMed
-
- Nelson CM, Jean RP, Tan JL, Liu WF, Sniadecki NJ, Spector AA, Chen CS,Proceedings of the National Academy of Sciences 102(33), 11594 (2005). DOI 10.1073/pnas.0502575102. URL http://www.pnas.org/cgi/doi/10.1073/pnas.0502575102 - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

