Application of an antibody chip for screening differentially expressed proteins during peach ripening and identification of a metabolon in the SAM cycle to generate a peach ethylene biosynthesis model
- PMID: 32194967
- PMCID: PMC7072073
- DOI: 10.1038/s41438-020-0249-9
Application of an antibody chip for screening differentially expressed proteins during peach ripening and identification of a metabolon in the SAM cycle to generate a peach ethylene biosynthesis model
Abstract
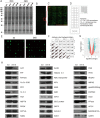
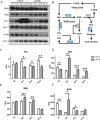
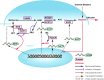
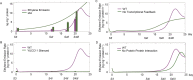
Peach (Prunus persica) is a typical climacteric fruit that produces ethylene rapidly during ripening, and its fruit softens quickly. Stony hard peach cultivars, however, do not produce large amounts of ethylene, and the fruit remains firm until fully ripe, thus differing from melting flesh peach cultivars. To identify the key proteins involved in peach fruit ripening, an antibody-based proteomic analysis was conducted. A mega-monoclonal antibody (mAb) library was generated and arrayed on a chip (mAbArray) at a high density, covering ~4950 different proteins of peach. Through the screening of peach fruit proteins with the mAbArray chip, differentially expressed proteins recognized by 1587 mAbs were identified, and 33 corresponding antigens were ultimately identified by immunoprecipitation and mass spectrometry. These proteins included not only important enzymes involved in ethylene biosynthesis, such as ACO1, SAHH, SAMS, and MetE, but also novel factors such as NUDT2. Furthermore, protein-protein interaction analysis identified a metabolon containing SAHH and MetE. By combining the antibody-based proteomic data with the transcriptomic and metabolic data, a mathematical model of ethylene biosynthesis in peach was constructed. Simulation results showed that MetE is an important regulator during peach ripening, partially through interaction with SAHH.
Keywords: Plant development; Protein-protein interaction networks.
© The Author(s) 2020.
Conflict of interest statement
Conflict of interestThe authors declare that they have no conflict of interest.
Figures


Similar articles
-
PpYUC11, a strong candidate gene for the stony hard phenotype in peach (Prunus persica L. Batsch), participates in IAA biosynthesis during fruit ripening.J Exp Bot. 2015 Dec;66(22):7031-44. doi: 10.1093/jxb/erv400. Epub 2015 Aug 24. J Exp Bot. 2015. PMID: 26307136 Free PMC article.
-
Genome-wide identification and transcriptome profiling reveal that E3 ubiquitin ligase genes relevant to ethylene, auxin and abscisic acid are differentially expressed in the fruits of melting flesh and stony hard peach varieties.BMC Genomics. 2019 Nov 21;20(1):892. doi: 10.1186/s12864-019-6258-0. BMC Genomics. 2019. PMID: 31752682 Free PMC article.
-
The involvement of 1-aminocyclopropane-1-carboxylic acid synthase isogene, Pp-ACS1, in peach fruit softening.J Exp Bot. 2006;57(6):1281-9. doi: 10.1093/jxb/erj097. Epub 2006 Mar 10. J Exp Bot. 2006. PMID: 16531466
-
Peach fruit ripening: A proteomic comparative analysis of the mesocarp of two cultivars with different flesh firmness at two ripening stages.Phytochemistry. 2011 Jul;72(10):1251-62. doi: 10.1016/j.phytochem.2011.01.012. Epub 2011 Feb 9. Phytochemistry. 2011. PMID: 21315381
-
Increased levels of IAA are required for system 2 ethylene synthesis causing fruit softening in peach (Prunus persica L. Batsch).J Exp Bot. 2013 Feb;64(4):1049-59. doi: 10.1093/jxb/ers381. Epub 2013 Jan 30. J Exp Bot. 2013. PMID: 23364941 Free PMC article.
Cited by
-
Shotgun proteomics of peach fruit reveals major metabolic pathways associated to ripening.BMC Genomics. 2021 Jan 6;22(1):17. doi: 10.1186/s12864-020-07299-y. BMC Genomics. 2021. PMID: 33413072 Free PMC article.
-
A Proteomics Insight into Advancements in the Rice-Microbe Interaction.Plants (Basel). 2023 Feb 28;12(5):1079. doi: 10.3390/plants12051079. Plants (Basel). 2023. PMID: 36903938 Free PMC article. Review.
-
Metabolons, enzyme-enzyme assemblies that mediate substrate channeling, and their roles in plant metabolism.Plant Commun. 2020 Jun 5;2(1):100081. doi: 10.1016/j.xplc.2020.100081. eCollection 2021 Jan 11. Plant Commun. 2020. PMID: 33511342 Free PMC article. Review.
-
The PPO family in Nicotiana tabacum is an important regulator to participate in pollination.BMC Plant Biol. 2024 Feb 9;24(1):102. doi: 10.1186/s12870-024-04769-3. BMC Plant Biol. 2024. PMID: 38331761 Free PMC article.
-
VvARF19 represses VvLBD13-mediated cell wall degradation to delay softening of grape berries.Hortic Res. 2024 Nov 18;12(2):uhae322. doi: 10.1093/hr/uhae322. eCollection 2025 Feb. Hortic Res. 2024. PMID: 40041604 Free PMC article.
References
-
- Layne, D. & Bassi, D. The Peach: Botany, Production & Uses (ed Layne, D.) 1–36 (CABI, Wallingford, 2008).
-
- Haji T, Yaegaki H, Yamaguchi M. Inheritance and expression of fruit texture melting, non-melting and stony hard in peach. Sci. Hortic. 2005;105:241–248. doi: 10.1016/j.scienta.2005.01.017. - DOI

