Emerging strategies to bridge the gap between pharmacogenomic research and its clinical implementation
- PMID: 32194983
- PMCID: PMC7057970
- DOI: 10.1038/s41525-020-0119-2
Emerging strategies to bridge the gap between pharmacogenomic research and its clinical implementation
Abstract
The genomic inter-individual heterogeneity remains a significant challenge for both clinical decision-making and the design of clinical trials. Although next-generation sequencing (NGS) is increasingly implemented in drug development and clinical trials, translation of the obtained genomic information into actionable clinical advice lags behind. Major reasons are the paucity of sufficiently powered trials that can quantify the added value of pharmacogenetic testing, and the considerable pharmacogenetic complexity with millions of rare variants with unclear functional consequences. The resulting uncertainty is reflected in inconsistencies of pharmacogenomic drug labels in Europe and the United States. In this review, we discuss how the knowledge gap for bridging pharmacogenomics into the clinics can be reduced. First, emerging methods that allow the high-throughput experimental characterization of pharmacogenomic variants combined with novel computational tools hold promise to improve the accuracy of drug response predictions. Second, tapping of large biobanks of therapeutic drug monitoring data allows to conduct high-powered retrospective studies that can validate the clinical importance of genetic variants, which are currently incompletely characterized. Combined, we are confident that these methods will improve the accuracy of drug response predictions and will narrow the gap between variant identification and its utilization for clinical decision-support.
Keywords: Genetic markers; Predictive markers.
© The Author(s) 2020.
Conflict of interest statement
Competing interestsV.M.L. and M.I.-S. are co-founders and shareholders of HepaPredict AB.
Figures


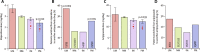
References
Publication types
LinkOut - more resources
Full Text Sources
Miscellaneous

