Compartmentalised acyl-CoA metabolism and roles in chromatin regulation
- PMID: 32199817
- PMCID: PMC7300382
- DOI: 10.1016/j.molmet.2020.01.005
Compartmentalised acyl-CoA metabolism and roles in chromatin regulation
Abstract
Background: Many metabolites serve as important signalling molecules to adjust cellular activities and functions based on nutrient availability. Links between acetyl-CoA metabolism, histone lysine acetylation, and gene expression have been documented and studied over the past decade. In recent years, several additional acyl modifications to histone lysine residues have been identified, which depend on acyl-coenzyme A thioesters (acyl-CoAs) as acyl donors. Acyl-CoAs are intermediates of multiple distinct metabolic pathways, and substantial evidence has emerged that histone acylation is metabolically sensitive. Nevertheless, the metabolic sources of acyl-CoAs used for chromatin modification in most cases remain poorly understood. Elucidating how these diverse chemical modifications are coupled to and regulated by cellular metabolism is important in deciphering their functional significance.
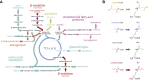
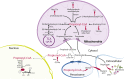
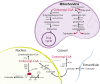
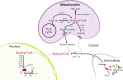
Scope of review: In this article, we review the metabolic pathways that produce acyl-CoAs, as well as emerging evidence for functional roles of diverse acyl-CoAs in chromatin regulation. Because acetyl-CoA has been extensively reviewed elsewhere, we will focus on four other acyl-CoA metabolites integral to major metabolic pathways that are also known to modify histones: succinyl-CoA, propionyl-CoA, crotonoyl-CoA, and butyryl-CoA. We also briefly mention several other acyl-CoA species, which present opportunities for further research; malonyl-CoA, glutaryl-CoA, 3-hydroxybutyryl-CoA, 2-hydroxyisobutyryl-CoA, and lactyl-CoA. Each acyl-CoA species has distinct roles in metabolism, indicating the potential to report shifts in the metabolic status of the cell. For each metabolite, we consider the metabolic pathways in which it participates and the nutrient sources from which it is derived, the compartmentalisation of its metabolism, and the factors reported to influence its abundance and potential nuclear availability. We also highlight reported biological functions of these metabolically-linked acylation marks. Finally, we aim to illuminate key questions in acyl-CoA metabolism as they relate to the control of chromatin modification.
Major conclusions: A majority of acyl-CoA species are annotated to mitochondrial metabolic processes. Since acyl-CoAs are not known to be directly transported across mitochondrial membranes, they must be synthesized outside of mitochondria and potentially within the nucleus to participate in chromatin regulation. Thus, subcellular metabolic compartmentalisation likely plays a key role in the regulation of histone acylation. Metabolite tracing in combination with targeting of relevant enzymes and transporters will help to map the metabolic pathways that connect acyl-CoA metabolism to chromatin modification. The specific function of each acyl-CoA may be determined in part by biochemical properties that affect its propensity for enzymatic versus non-enzymatic protein modification, as well as the various enzymes that can add, remove and bind each modification. Further, competitive and inhibitory effects of different acyl-CoA species on these enzymes make determining the relative abundance of acyl-CoA species in specific contexts important to understand the regulation of chromatin acylation. An improved and more nuanced understanding of metabolic regulation of chromatin and its roles in physiological and disease-related processes will emerge as these questions are answered.
Keywords: Acyl-CoA; Acylation; Compartmentalisation; Histone; Metabolism.
Copyright © 2020 The Author(s). Published by Elsevier GmbH.. All rights reserved.
Figures

References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

