Metabolomics hallmarks OPA1 variants correlating with their in vitro phenotype and predicting clinical severity
- PMID: 32202296
- PMCID: PMC7254852
- DOI: 10.1093/hmg/ddaa047
Metabolomics hallmarks OPA1 variants correlating with their in vitro phenotype and predicting clinical severity
Abstract
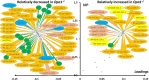
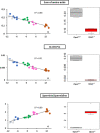
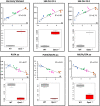
Interpretation of variants of uncertain significance is an actual major challenge. We addressed this question on a set of OPA1 missense variants responsible for variable severity of neurological impairments. We used targeted metabolomics to explore the different signatures of OPA1 variants expressed in Opa1 deleted mouse embryonic fibroblasts (Opa1-/- MEFs), grown under selective conditions. Multivariate analyses of data discriminated Opa1+/+ from Opa1-/- MEFs metabolic signatures and classified OPA1 variants according to their in vitro severity. Indeed, the mild p.I382M hypomorphic variant was segregating close to the wild-type allele, while the most severe p.R445H variant was close to Opa1-/- MEFs, and the p.D603H and p.G439V alleles, responsible for isolated and syndromic presentations, respectively, were intermediary between the p.I382M and the p.R445H variants. The most discriminant metabolic features were hydroxyproline, the spermine/spermidine ratio, amino acid pool and several phospholipids, emphasizing proteostasis, endoplasmic reticulum (ER) stress and phospholipid remodeling as the main mechanisms ranking OPA1 allele impacts on metabolism. These results demonstrate the high resolving power of metabolomics in hierarchizing OPA1 missense mutations by their in vitro severity, fitting clinical expressivity. This suggests that our methodological approach can be used to discriminate the pathological significance of variants in genes responsible for other rare metabolic diseases and may be instrumental to select possible compounds eligible for supplementation treatment.
© The Author(s) 2020. Published by Oxford University Press. All rights reserved. For Permissions, please email: journals.permissions@oup.com.
Figures


Similar articles
-
Mitochondrial dysfunction in an Opa1(Q285STOP) mouse model of dominant optic atrophy results from Opa1 haploinsufficiency.Cell Death Dis. 2016 Jul 28;7(7):e2309. doi: 10.1038/cddis.2016.160. Cell Death Dis. 2016. PMID: 27468686 Free PMC article.
-
Deciphering OPA1 mutations pathogenicity by combined analysis of human, mouse and yeast cell models.Biochim Biophys Acta Mol Basis Dis. 2018 Oct;1864(10):3496-3514. doi: 10.1016/j.bbadis.2018.08.004. Epub 2018 Aug 4. Biochim Biophys Acta Mol Basis Dis. 2018. PMID: 30293569
-
The Metabolomic Signature of Opa1 Deficiency in Rat Primary Cortical Neurons Shows Aspartate/Glutamate Depletion and Phospholipids Remodeling.Sci Rep. 2019 Apr 15;9(1):6107. doi: 10.1038/s41598-019-42554-7. Sci Rep. 2019. PMID: 30988455 Free PMC article.
-
OPA1: How much do we know to approach therapy?Pharmacol Res. 2018 May;131:199-210. doi: 10.1016/j.phrs.2018.02.018. Epub 2018 Feb 15. Pharmacol Res. 2018. PMID: 29454676 Review.
-
OPA1 (Kjer type) dominant optic atrophy: a novel mitochondrial disease.Mol Genet Metab. 2002 Feb;75(2):97-107. doi: 10.1006/mgme.2001.3278. Mol Genet Metab. 2002. PMID: 11855928 Review.
Cited by
-
Dominant Optic Atrophy (DOA): Modeling the Kaleidoscopic Roles of OPA1 in Mitochondrial Homeostasis.Front Neurol. 2021 Jun 9;12:681326. doi: 10.3389/fneur.2021.681326. eCollection 2021. Front Neurol. 2021. PMID: 34177786 Free PMC article. Review.
-
3D reconstruction of murine mitochondria reveals changes in structure during aging linked to the MICOS complex.Aging Cell. 2023 Dec;22(12):e14009. doi: 10.1111/acel.14009. Epub 2023 Nov 13. Aging Cell. 2023. PMID: 37960952 Free PMC article.
-
Mutations at a split codon in the GTPase-encoding domain of OPA1 cause dominant optic atrophy through different molecular mechanisms.Hum Mol Genet. 2022 Mar 3;31(5):761-774. doi: 10.1093/hmg/ddab286. Hum Mol Genet. 2022. PMID: 34559197 Free PMC article.
-
Metabolic reprogramming in the OPA1-deficient cells.Cell Mol Life Sci. 2022 Sep 14;79(10):517. doi: 10.1007/s00018-022-04542-5. Cell Mol Life Sci. 2022. PMID: 36103091 Free PMC article.
-
The loss of OPA1 accelerates intervertebral disc degeneration and osteoarthritis in aged mice.Nat Commun. 2025 Jul 1;16(1):5996. doi: 10.1038/s41467-025-60933-9. Nat Commun. 2025. PMID: 40595596 Free PMC article.
References
-
- Ferré M., Caignard A., Milea D., Leruez S., Cassereau J., Chevrollier A., Amati-Bonneau P., Verny C., Bonneau D., Procaccio V. and Reynier P. (2015) Improved locus-specific database for OPA1 mutations allows inclusion of advanced clinical data. Hum. Mutat., 36, 20–25. - PubMed
-
- Olichon A., Baricault L., Gas N., Guillou E., Valette A., Belenguer P. and Lenaers G. (2003) Loss of OPA1 perturbates the mitochondrial inner membrane structure and integrity, leading to cytochrome c release and apoptosis. J. Biol. Chem., 278, 7743–7746. - PubMed
-
- Olichon A., Elachouri G., Baricault L., Delettre C., Belenguer P. and Lenaers G. (2007) OPA1 alternate splicing uncouples an evolutionary conserved function in mitochondrial fusion from a vertebrate restricted function in apoptosis. Cell Death Differ., 14, 682–692. - PubMed
-
- Zanna C., Ghelli A., Porcelli A.M., Karbowski M., Youle R.J., Schimpf S., Wissinger B., Pinti M., Cossarizza A., Vidoni S. et al. (2008) OPA1 mutations associated with dominant optic atrophy impair oxidative phosphorylation and mitochondrial fusion. Brain, 131, 352–367. - PubMed

