Three-dimensional chromatin landscapes in T cell acute lymphoblastic leukemia
- PMID: 32203470
- PMCID: PMC7138649
- DOI: 10.1038/s41588-020-0602-9
Three-dimensional chromatin landscapes in T cell acute lymphoblastic leukemia
Abstract
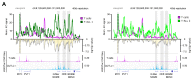
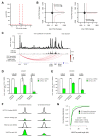
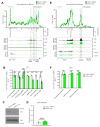
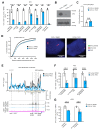
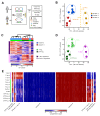
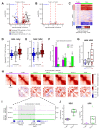
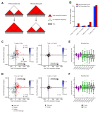
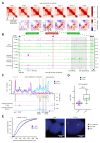
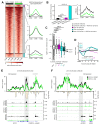
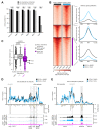
Differences in three-dimensional (3D) chromatin architecture can influence the integrity of topologically associating domains (TADs) and rewire specific enhancer-promoter interactions, impacting gene expression and leading to human disease. Here we investigate the 3D chromatin architecture in T cell acute lymphoblastic leukemia (T-ALL) by using primary human leukemia specimens and examine the dynamic responses of this architecture to pharmacological agents. Systematic integration of matched in situ Hi-C, RNA-seq and CTCF ChIP-seq datasets revealed widespread differences in intra-TAD chromatin interactions and TAD boundary insulation in T-ALL. Our studies identify and focus on a TAD 'fusion' event associated with absence of CTCF-mediated insulation, enabling direct interactions between the MYC promoter and a distal super-enhancer. Moreover, our data also demonstrate that small-molecule inhibitors targeting either oncogenic signal transduction or epigenetic regulation can alter specific 3D interactions found in leukemia. Overall, our study highlights the impact, complexity and dynamic nature of 3D chromatin architecture in human acute leukemia.
Conflict of interest statement
A.T. is a Scientific Advisor to Intelligencia.AI. All other authors declare that they have no competing financial interests.
Figures
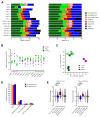
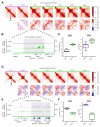
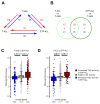
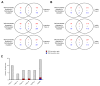


References
-
- Spielmann M, Lupianez DG, Mundlos S. Structural variation in the 3D genome. Nat Rev Genet. 2018 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- R01 CA133379/CA/NCI NIH HHS/United States
- R01 CA194923/CA/NCI NIH HHS/United States
- R01 CA228135/CA/NCI NIH HHS/United States
- R01 CA149655/CA/NCI NIH HHS/United States
- R01 GM127538/GM/NIGMS NIH HHS/United States
- R01 CA216421/CA/NCI NIH HHS/United States
- R01 CA202025/CA/NCI NIH HHS/United States
- UG1 CA233332/CA/NCI NIH HHS/United States
- U54 CA193419/CA/NCI NIH HHS/United States
- R35 CA220499/CA/NCI NIH HHS/United States
- P01 CA229086/CA/NCI NIH HHS/United States
- P30 CA016087/CA/NCI NIH HHS/United States
- 639784/ERC_/European Research Council/International
- U10 CA180820/CA/NCI NIH HHS/United States
- R00 CA188293/CA/NCI NIH HHS/United States
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

