Transcriptome reconstruction and functional analysis of eukaryotic marine plankton communities via high-throughput metagenomics and metatranscriptomics
- PMID: 32205368
- PMCID: PMC7197479
- DOI: 10.1101/gr.253070.119
Transcriptome reconstruction and functional analysis of eukaryotic marine plankton communities via high-throughput metagenomics and metatranscriptomics
Abstract
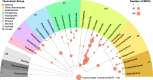
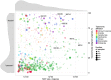
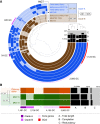
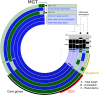
Large-scale metagenomic and metatranscriptomic data analyses are often restricted by their gene-centric approach, limiting the ability to understand organismal and community biology. De novo assembly of large and mosaic eukaryotic genomes from complex meta-omics data remains a challenging task, especially in comparison with more straightforward bacterial and archaeal systems. Here, we use a transcriptome reconstruction method based on clustering co-abundant genes across a series of metagenomic samples. We investigated the co-abundance patterns of ∼37 million eukaryotic unigenes across 365 metagenomic samples collected during the Tara Oceans expeditions to assess the diversity and functional profiles of marine plankton. We identified ∼12,000 co-abundant gene groups (CAGs), encompassing ∼7 million unigenes, including 924 metagenomics-based transcriptomes (MGTs, CAGs larger than 500 unigenes). We demonstrated the biological validity of the MGT collection by comparing individual MGTs with available references. We identified several key eukaryotic organisms involved in dimethylsulfoniopropionate (DMSP) biosynthesis and catabolism in different oceanic provinces, thus demonstrating the potential of the MGT collection to provide functional insights on eukaryotic plankton. We established the ability of the MGT approach to capture interspecies associations through the analysis of a nitrogen-fixing haptophyte-cyanobacterial symbiotic association. This MGT collection provides a valuable resource for analyses of eukaryotic plankton in the open ocean by giving access to the genomic content and functional potential of many ecologically relevant eukaryotic species.
© 2020 Vorobev et al.; Published by Cold Spring Harbor Laboratory Press.
Figures

References
-
- Armbrecht LH, Eriksen R, Leventer A, Armand LK. 2017. First observations of living sea-ice diatom agglomeration to tintinnid loricae in East Antarctica. J Plankton Res 39: 795–802. 10.1093/plankt/fbx036 - DOI
-
- Ayers GP, Cainey JM. 2007. The CLAW hypothesis: a review of the major developments. Environ Chem 4: 366–374. 10.1071/EN07080 - DOI
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
