Identification of slit3 as a locus affecting nicotine preference in zebrafish and human smoking behaviour
- PMID: 32209227
- PMCID: PMC7096180
- DOI: 10.7554/eLife.51295
Identification of slit3 as a locus affecting nicotine preference in zebrafish and human smoking behaviour
Abstract
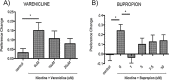
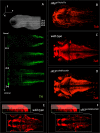
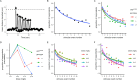
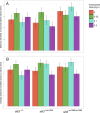
To facilitate smoking genetics research we determined whether a screen of mutagenized zebrafish for nicotine preference could predict loci affecting smoking behaviour. From 30 screened F3 sibling groups, where each was derived from an individual ethyl-nitrosurea mutagenized F0 fish, two showed increased or decreased nicotine preference. Out of 25 inactivating mutations carried by the F3 fish, one in the slit3 gene segregated with increased nicotine preference in heterozygous individuals. Focussed SNP analysis of the human SLIT3 locus in cohorts from UK (n=863) and Finland (n=1715) identified two variants associated with cigarette consumption and likelihood of cessation. Characterisation of slit3 mutant larvae and adult fish revealed decreased sensitivity to the dopaminergic and serotonergic antagonist amisulpride, known to affect startle reflex that is correlated with addiction in humans, and increased htr1aa mRNA expression in mutant larvae. No effect on neuronal pathfinding was detected. These findings reveal a role for SLIT3 in development of pathways affecting responses to nicotine in zebrafish and smoking in humans.
Keywords: acoustic startle; conditioned place preference; gene association; neuroscience; nicotine; slit3; smoking; zebrafish.
© 2020, García-González et al.
Conflict of interest statement
JG, AB, MP, RR, DJ, AS, MT, EB, DS, AM, JK, TP, VK, RW, CB No competing interests declared
Figures








Similar articles
-
Methionine Supplementation Abolishes Nicotine-Induced Place Preference in Zebrafish: a Behavioral and Molecular Analysis.Mol Neurobiol. 2021 Jun;58(6):2590-2607. doi: 10.1007/s12035-020-02260-2. Epub 2021 Jan 21. Mol Neurobiol. 2021. PMID: 33475949
-
Evaluation of the rewarding properties of nicotine and caffeine by implementation of a five-choice conditioned place preference task in zebrafish.Prog Neuropsychopharmacol Biol Psychiatry. 2018 Jun 8;84(Pt A):160-172. doi: 10.1016/j.pnpbp.2018.02.001. Epub 2018 Feb 23. Prog Neuropsychopharmacol Biol Psychiatry. 2018. PMID: 29481898
-
Dissociation of food and opiate preference by a genetic mutation in zebrafish.Genes Brain Behav. 2006 Oct;5(7):497-505. doi: 10.1111/j.1601-183X.2005.00185.x. Genes Brain Behav. 2006. PMID: 17010096
-
Neurogenetic determinants and mechanisms of addiction to nicotine and smoked tobacco.Eur J Neurosci. 2019 Aug;50(3):2164-2179. doi: 10.1111/ejn.14171. Epub 2018 Oct 25. Eur J Neurosci. 2019. PMID: 30256469 Review.
-
Inter-relationships between conditioned and primary reinforcement in the maintenance of cigarette smoking.Br J Addict. 1991 May;86(5):605-9. doi: 10.1111/j.1360-0443.1991.tb01816.x. Br J Addict. 1991. PMID: 1859927 Review.
Cited by
-
Identification, conservation, and expression of tiered pharmacogenes in zebrafish.PLoS One. 2022 Aug 30;17(8):e0273582. doi: 10.1371/journal.pone.0273582. eCollection 2022. PLoS One. 2022. PMID: 36040978 Free PMC article.
-
Identification of nicotine-seeking and avoiding larval zebrafish using a new three-choice behavioral assay.Front Mol Neurosci. 2023 Mar 30;16:1112927. doi: 10.3389/fnmol.2023.1112927. eCollection 2023. Front Mol Neurosci. 2023. PMID: 37063370 Free PMC article.
-
Integration of cooperative and opposing molecular programs drives learning-associated behavioral plasticity.PLoS Genet. 2023 Mar 27;19(3):e1010650. doi: 10.1371/journal.pgen.1010650. eCollection 2023 Mar. PLoS Genet. 2023. PMID: 36972301 Free PMC article.
-
Methionine Supplementation Abolishes Nicotine-Induced Place Preference in Zebrafish: a Behavioral and Molecular Analysis.Mol Neurobiol. 2021 Jun;58(6):2590-2607. doi: 10.1007/s12035-020-02260-2. Epub 2021 Jan 21. Mol Neurobiol. 2021. PMID: 33475949
-
Novel DNA methylation changes in mouse lungs associated with chronic smoking.Epigenetics. 2024 Dec;19(1):2322386. doi: 10.1080/15592294.2024.2322386. Epub 2024 Mar 4. Epigenetics. 2024. PMID: 38436597 Free PMC article.
References
-
- Ackerman KM, Boyd RT. Nicotinic Acetylcholine Receptor Technologies. New York: Springer; 2016. Analysis of Nicotinic Acetylcholine Receptor (nAChR) Gene Expression in Zebrafish (Danio rerio) by In Situ Hybridization and PCR; pp. 1–31. - DOI
Publication types
MeSH terms
Substances
Grants and funding
- R01 AA015325/AA/NIAAA NIH HHS/United States
- G1000053/NC3RS_/National Centre for the Replacement, Refinement and Reduction of Animals in Research/United Kingdom
- U01 DA044400/DA/NIDA NIH HHS/United States
- BB/M007863/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- G1000403/MRC_/Medical Research Council/United Kingdom
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

