Role of changes in SARS-CoV-2 spike protein in the interaction with the human ACE2 receptor: An in silico analysis
- PMID: 32210742
- PMCID: PMC7081066
- DOI: 10.17179/excli2020-1167
Role of changes in SARS-CoV-2 spike protein in the interaction with the human ACE2 receptor: An in silico analysis
Abstract
Many human viral diseases are a consequence of a zoonotic event. Some of the diseases caused by these zoonotic events have affected millions of people around the world, some of which have resulted in high rates of morbidity/mortality in humans. Changes in the viral proteins that function as ligands of the host receptor may promote the spillover between species. The most recent of these zoonotic events that have caused an ongoing epidemic of high magnitude is the Covid-19 epidemics caused by SARS-CoV-2. The aim of this study was to determine the mutation(s) in the sequence of the spike protein of the SARS-CoV-2 that might be favoring human to human transmission. An in silico approach was performed, and changes were detected in the S1 subunit of the receptor-binding domain of spike. The observed changes have significant effect on SARS-CoV-2 spike/ACE2 interaction and produce a reduction in the binding energy, compared to the one of the Bat-CoV to this receptor. The data presented in this study suggest a higher affinity of the SARS-Cov-2 spike protein to the human ACE2 receptor, compared to the one of Bat-CoV spike and ACE2. This could be the cause of the rapid viral spread of SARS-CoV-2 in humans.
Keywords: ACE2; Coronavirus; SARS-CoV-2; Spike; outbreak.
Copyright © 2020 Ortega et al.
Figures



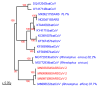

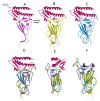
References
-
- Arnold K, Bordoli L, Kopp J, Schwede T. The SWISS-MODEL workspace: A web-based environment for protein structure homology modelling. Bioinformatics. 2006;22:195–201. - PubMed
-
- Laskowski RA, MacArthur MW, Moss DS, Thornton JM. PROCHECK - a program to check the stereochemical quality of protein structures. J Appl Crystallogr. 1993;26:283–91.
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous
