Whole Genome Sequencing Differentiates Presumptive Extended Spectrum Beta-Lactamase Producing Escherichia coli along Segments of the One Health Continuum
- PMID: 32235751
- PMCID: PMC7143971
- DOI: 10.3390/microorganisms8030448
Whole Genome Sequencing Differentiates Presumptive Extended Spectrum Beta-Lactamase Producing Escherichia coli along Segments of the One Health Continuum
Abstract
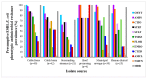
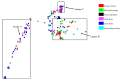
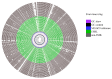
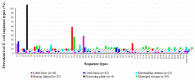
Antimicrobial resistance (AMR) has important implications for the continued use of antibiotics to control infectious diseases in both beef cattle and humans. AMR along the One Health continuum of the beef production system is largely unknown. Here, whole genomes of presumptive extended-spectrum β-lactamase E. coli (ESBL-EC) from cattle feces (n = 40), feedlot catch basins (n = 42), surrounding streams (n = 21), a beef processing plant (n = 4), municipal sewage (n = 30), and clinical patients (n = 25) are described. ESBL-EC were isolated from ceftriaxone selective plates and subcultured on ampicillin selective plates. Agreement of genotype-phenotype prediction of AMR ranged from 93.2% for ampicillin to 100% for neomycin, trimethoprim/sulfamethoxazole, and enrofloxacin resistance. Overall, β-lactam (100%; blaEC, blaTEM-1, blaSHV, blaOXA, blaCTX-M-), tetracycline (90.1%; tet(A), tet(B)) and folate synthesis (sul2) antimicrobial resistance genes (ARGs) were most prevalent. The ARGs tet(C), tet(M), tet(32), blaCTX-M-1, blaCTX-M-14, blaOXA-1, dfrA18, dfrA19, catB3, and catB4 were exclusive to human sources, while blaTEM-150, blaSHV-11-12, dfrA12, cmlA1, and cmlA5 were exclusive to beef cattle sources. Frequently encountered virulence factors across all sources included adhesion and type II and III secretion systems, while IncFIB(AP001918) and IncFII plasmids were also common. Specificity and prevalence of ARGs between cattle-sourced and human-sourced presumptive ESBL-EC likely reflect differences in antimicrobial use in cattle and humans. Comparative genomics revealed phylogenetically distinct clusters for isolates from human vs. cattle sources, implying that human infections caused by ESBL-EC in this region might not originate from beef production sources.
Keywords: One Health; antimicrobial resistance; comparative genomics; extended spectrum Beta-lactamase producing E. coli; whole genome sequencing.
Conflict of interest statement
CWB is part owner and managing partner of Feedlot Health Management Services and Southern Alberta Veterinary Services. SJH is an employee at Feedlot Health Management Services, Okotoks, Alberta, Canada. Feedlot Health is a private company that provides expert consultation regarding management and production of calf grower calves and feedlot cattle, including developing veterinary protocols to support animal health. Feedlot Health also conducts in-house and contract research related to dairy calf grower and feedlot production.
Figures



References
-
- Johnson J.R., Nicolas-Chanoine M.H., DebRoy C., Castanheira M., Robicsek A., Hansen G., Weissman S., Urban C., Platell J., Trott D., et al. Comparison of Escherichia coli ST131 pulsotypes, by epidemiologic traits, 1967–2009. Emerg. Infect. Dis. 2012;18:598–607. doi: 10.3201/eid1804.111627. - DOI - PMC - PubMed
-
- World Health Organization [(accessed on 27 February 2017)];Global Priority List of Antibiotic-Resistant Bacteria to Guide Research, Discovery, and Development of New Antibiotics, WHO Essential Medicines and Health Products Guidelines. 2017 Available online: https://www.who.int/medicines/publications/global-priority-list-antibiot...
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

