CRISPR Tools To Control Gene Expression in Bacteria
- PMID: 32238445
- PMCID: PMC7117552
- DOI: 10.1128/MMBR.00077-19
CRISPR Tools To Control Gene Expression in Bacteria
Abstract
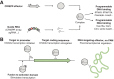
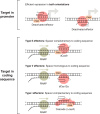
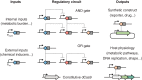
CRISPR-Cas systems have been engineered as powerful tools to control gene expression in bacteria. The most common strategy relies on the use of Cas effectors modified to bind target DNA without introducing DNA breaks. These effectors can either block the RNA polymerase or recruit it through activation domains. Here, we discuss the mechanistic details of how Cas effectors can modulate gene expression by blocking transcription initiation or acting as transcription roadblocks. CRISPR-Cas tools can be further engineered to obtain fine-tuned control of gene expression or target multiple genes simultaneously. Several caveats in using these tools have also been revealed, including off-target effects and toxicity, making it important to understand the design rules of engineered CRISPR-Cas effectors in bacteria. Alternatively, some types of CRISPR-Cas systems target RNA and could be used to block gene expression at the posttranscriptional level. Finally, we review applications of these tools in high-throughput screens and the progress and challenges in introducing CRISPR knockdown to other species, including nonmodel bacteria with industrial or clinical relevance. A deep understanding of how CRISPR-Cas systems can be harnessed to control gene expression in bacteria and build powerful tools will certainly open novel research directions.
Keywords: CRISPR; gene silencing; transcriptional regulation.
Copyright © 2020 American Society for Microbiology.
Figures
References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

