A Metformin-Responsive Metabolic Pathway Controls Distinct Steps in Gastric Progenitor Fate Decisions and Maturation
- PMID: 32243780
- PMCID: PMC7275895
- DOI: 10.1016/j.stem.2020.03.006
A Metformin-Responsive Metabolic Pathway Controls Distinct Steps in Gastric Progenitor Fate Decisions and Maturation
Abstract
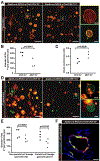
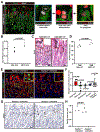
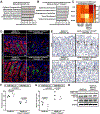
Cellular metabolism plays important functions in dictating stem cell behaviors, although its role in stomach epithelial homeostasis has not been evaluated in depth. Here, we show that the energy sensor AMP kinase (AMPK) governs gastric epithelial progenitor differentiation. Administering the AMPK activator metformin decreases epithelial progenitor proliferation and increases acid-secreting parietal cells (PCs) in mice and organoids. AMPK activation targets Krüppel-like factor 4 (KLF4), known to govern progenitor proliferation and PC fate choice, and PGC1α, which we show controls PC maturation after their specification. PC-specific deletion of AMPKα or PGC1α causes defective PC maturation, which could not be rescued by metformin. However, metformin treatment still increases KLF4 levels and suppresses progenitor proliferation. Thus, AMPK activates KLF4 in progenitors to reduce self-renewal and promote PC fate, whereas AMPK-PGC1α activation within the PC lineage promotes maturation, providing a potential suggestion for why metformin increases acid secretion and reduces gastric cancer risk in humans.
Keywords: ADM; SPEM; mTORC1; metaplasia; mitochondria; paligenosis; ribosomes.
Copyright © 2020 Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of Interests The authors declare no competing interests.
Figures




References
-
- Abe K, Irie K, Nakanishi H, Suzuki H, and Fujiyoshi Y (2018). Crystal structures of the gastric proton pump. Nature 556, 214–218. - PubMed
-
- Adami JG (1900). On growth and overgrowth In “Festschrift” in honor of Abraham Jacobi, MD, LLD: to commemorate the seventieth anniversary of his birth, May sixth, 1900, Huber F, Sondern FE (eds) (New Rochelle, NY: Knickerbocker Press; ), pp. 422–432.
-
- Adami JG, and Nicholls AG (1910). The principles of pathology (Philadelphia: : Lea & Febiger, 1908-1909).
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

