Transcriptome Analysis Reveals the Neuro-Immune Interactions in Duck Tembusu Virus-Infected Brain
- PMID: 32244328
- PMCID: PMC7177238
- DOI: 10.3390/ijms21072402
Transcriptome Analysis Reveals the Neuro-Immune Interactions in Duck Tembusu Virus-Infected Brain
Abstract
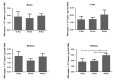
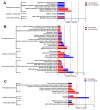
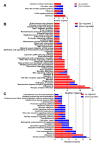
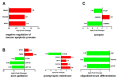
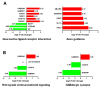
The duck Tembusu virus (DTMUV) is a mosquito-borne flavivirus. It causes severe symptoms of egg-drop, as well as neurological symptoms and brain damage in ducks. However, the specific molecular mechanisms of DTMUV-induced neurovirulence and host responses in the brain remain obscure. To better understand the host-pathogen and neuro-immune interactions of DTMUV infection, we conducted high-throughput RNA-sequencing to reveal the transcriptome profiles of DTMUV-infected duck brain. Totals of 117, 212, and 150 differentially expressed genes (DEGs) were identified at 12, 24, and 48 h post infection (hpi). Gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analyses uncovered genes and pathways related to the nervous system and immune responses in duck brain. Neuro-related genes, including WNT3A, GATA3, and CHRNA6, were found to be significantly downregulated. RIG-I-like receptors (DHX58, IFIH1) and Toll-like receptors (TLR2 and TLR3) were activated, inducing the expression of 22 interferon stimulated genes (ISGs) and antigen-processing and -presenting genes (TAP1 and TAP2) in the brain. Our research provides comprehensive information for the molecular mechanisms of neuro-immune and host-pathogen interactions of DTMUV.
Keywords: DTMUV; Transcriptome analysis; brain; host–pathogen interaction; molecular mechanism; neuro-immune interaction.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures



Similar articles
-
Differently Expression Analysis and Function Prediction of Long Non-coding RNAs in Duck Embryo Fibroblast Cells Infected by Duck Tembusu Virus.Front Immunol. 2020 Aug 4;11:1729. doi: 10.3389/fimmu.2020.01729. eCollection 2020. Front Immunol. 2020. PMID: 32849615 Free PMC article.
-
Comparative Transcriptomic Analysis of Immune-Related Gene Expression in Duck Embryo Fibroblasts Following Duck Tembusu Virus Infection.Int J Mol Sci. 2018 Aug 8;19(8):2328. doi: 10.3390/ijms19082328. Int J Mol Sci. 2018. PMID: 30096804 Free PMC article.
-
Transcriptome analysis reveals new insight of duck Tembusu virus (DTMUV)-infected DF-1 cells.Res Vet Sci. 2021 Jul;137:150-158. doi: 10.1016/j.rvsc.2021.04.028. Epub 2021 Apr 29. Res Vet Sci. 2021. PMID: 33975194
-
Innate immune responses to duck Tembusu virus infection.Vet Res. 2020 Jul 8;51(1):87. doi: 10.1186/s13567-020-00814-9. Vet Res. 2020. PMID: 32641107 Free PMC article. Review.
-
Advancements in Research on Duck Tembusu Virus Infections.Viruses. 2024 May 20;16(5):811. doi: 10.3390/v16050811. Viruses. 2024. PMID: 38793692 Free PMC article. Review.
Cited by
-
Molecular Research on Vector-Borne Diseases of Medical Interest: From Bench to Application 2.0.Int J Mol Sci. 2023 Apr 26;24(9):7907. doi: 10.3390/ijms24097907. Int J Mol Sci. 2023. PMID: 37175612 Free PMC article.
-
Differently Expression Analysis and Function Prediction of Long Non-coding RNAs in Duck Embryo Fibroblast Cells Infected by Duck Tembusu Virus.Front Immunol. 2020 Aug 4;11:1729. doi: 10.3389/fimmu.2020.01729. eCollection 2020. Front Immunol. 2020. PMID: 32849615 Free PMC article.
-
Duck LGP2 Downregulates RIG-I Signaling Pathway-Mediated Innate Immunity Against Tembusu Virus.Front Immunol. 2022 Jun 15;13:916350. doi: 10.3389/fimmu.2022.916350. eCollection 2022. Front Immunol. 2022. PMID: 35784309 Free PMC article.
-
Duck Tembusu virus induces incomplete autophagy via the ERK/mTOR and AMPK/mTOR signalling pathways to promote viral replication in neuronal cells.Vet Res. 2023 Nov 7;54(1):103. doi: 10.1186/s13567-023-01235-0. Vet Res. 2023. PMID: 37936178 Free PMC article.
-
Differential gene expression reveals host factors for viral shedding variation in mallards (Anas platyrhynchos) infected with low-pathogenic avian influenza virus.J Gen Virol. 2022 Mar;103(3):10.1099/jgv.0.001724. doi: 10.1099/jgv.0.001724. J Gen Virol. 2022. PMID: 35353676 Free PMC article.
References
MeSH terms
Substances
Supplementary concepts
Grants and funding
- SQZS1802/Guangdong province key laboratory of waterfowl healthy breeding open project
- 2019KJ137/Modern agricultural industry technology system innovation team of Guangdong Province
- R2018QD-093/Special fund for scientific innovation strategy-construction of high level Academy of Agriculture Science
- 2017A040403015/Science and Technology Planning Project of Guangdong Province
- 2019B020218004, 2019B020217002/Key-Area Research and Development Program of Guangdong Province
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

